+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Shaker in low K+ (4mM K+) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | voltage-gated potassium channel / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmating behavior, sex discrimination / Phase 3 - rapid repolarisation / behavioral response to ether / Voltage gated Potassium channels / proboscis extension reflex / larval locomotory behavior / regulation of synaptic activity / courtship behavior / positive regulation of membrane potential / positive regulation of circadian sleep/wake cycle, sleep ...mating behavior, sex discrimination / Phase 3 - rapid repolarisation / behavioral response to ether / Voltage gated Potassium channels / proboscis extension reflex / larval locomotory behavior / regulation of synaptic activity / courtship behavior / positive regulation of membrane potential / positive regulation of circadian sleep/wake cycle, sleep / regulation of circadian sleep/wake cycle, sleep / detection of visible light / voltage-gated monoatomic cation channel activity / sleep / cellular response to dopamine / delayed rectifier potassium channel activity / axon extension / action potential / voltage-gated potassium channel activity / potassium ion transmembrane transport / voltage-gated potassium channel complex / potassium ion transport / protein homooligomerization / sensory perception of taste / perikaryon / learning or memory / neuron projection / membrane raft / membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.39 Å | |||||||||
 Authors Authors | Tan X / Swartz KJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2023 Journal: Sci Adv / Year: 2023Title: Eukaryotic Kv channel Shaker inactivates through selectivity filter dilation rather than collapse. Authors: Robyn Stix / Xiao-Feng Tan / Chanhyung Bae / Ana I Fernández-Mariño / Kenton J Swartz / José D Faraldo-Gómez /  Abstract: Eukaryotic voltage-gated K channels have been extensively studied, but the structural bases for some of their most salient functional features remain to be established. C-type inactivation, for ...Eukaryotic voltage-gated K channels have been extensively studied, but the structural bases for some of their most salient functional features remain to be established. C-type inactivation, for example, is an auto-inhibitory mechanism that confers temporal resolution to their signal-firing activity. In a recent breakthrough, studies of a mutant of Shaker that is prone to inactivate indicated that this process entails a dilation of the selectivity filter, the narrowest part of the ion conduction pathway. Here, we report an atomic-resolution cryo-electron microscopy structure that demonstrates that the wild-type channel can also adopt this dilated state. All-atom simulations corroborate this conformation is congruent with the electrophysiological characteristics of the C-type inactivated state, namely, residual K conductance and altered ion specificity, and help rationalize why inactivation is accelerated or impeded by certain mutations. In summary, this study establishes the molecular basis for an important self-regulatory mechanism in eukaryotic K channels, laying a solid foundation for further studies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41193.map.gz emd_41193.map.gz | 95.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41193-v30.xml emd-41193-v30.xml emd-41193.xml emd-41193.xml | 15 KB 15 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_41193.png emd_41193.png | 39.5 KB | ||
| Filedesc metadata |  emd-41193.cif.gz emd-41193.cif.gz | 5.8 KB | ||
| Others |  emd_41193_half_map_1.map.gz emd_41193_half_map_1.map.gz emd_41193_half_map_2.map.gz emd_41193_half_map_2.map.gz | 76 MB 76 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41193 http://ftp.pdbj.org/pub/emdb/structures/EMD-41193 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41193 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41193 | HTTPS FTP |
-Validation report
| Summary document |  emd_41193_validation.pdf.gz emd_41193_validation.pdf.gz | 757.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41193_full_validation.pdf.gz emd_41193_full_validation.pdf.gz | 757.4 KB | Display | |
| Data in XML |  emd_41193_validation.xml.gz emd_41193_validation.xml.gz | 12.6 KB | Display | |
| Data in CIF |  emd_41193_validation.cif.gz emd_41193_validation.cif.gz | 14.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41193 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41193 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41193 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41193 | HTTPS FTP |
-Related structure data
| Related structure data |  8teoMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41193.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41193.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_41193_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
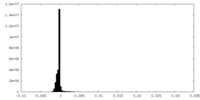
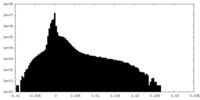
| Density Histograms |
-Half map: #2
| File | emd_41193_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
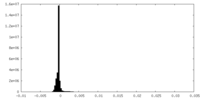
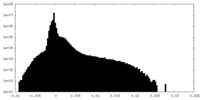
| Density Histograms |
- Sample components
Sample components
-Entire : voltage-gated potassium channel Shaker
| Entire | Name: voltage-gated potassium channel Shaker |
|---|---|
| Components |
|
-Supramolecule #1: voltage-gated potassium channel Shaker
| Supramolecule | Name: voltage-gated potassium channel Shaker / type: cell / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Potassium voltage-gated channel protein Shaker
| Macromolecule | Name: Potassium voltage-gated channel protein Shaker / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 74.525953 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GTMAAVAGLY GLGEDRQHRK KQQQQQQHQK EQLEQKEEQK KIAERKLQLR EQQLQRNSLD GYGSLPKLSS QDEEGGAGHG FGGGPQHFE PIPHDHDFCE RVVINVSGLR FETQLRTLNQ FPDTLLGDPA RRLRYFDPLR NEYFFDRSRP SFDAILYYYQ S GGRLRRPV ...String: GTMAAVAGLY GLGEDRQHRK KQQQQQQHQK EQLEQKEEQK KIAERKLQLR EQQLQRNSLD GYGSLPKLSS QDEEGGAGHG FGGGPQHFE PIPHDHDFCE RVVINVSGLR FETQLRTLNQ FPDTLLGDPA RRLRYFDPLR NEYFFDRSRP SFDAILYYYQ S GGRLRRPV NVPLDVFSEE IKFYELGDQA INKFREDEGF IKEEERPLPD NEKQRKVWLL FEYPESSQAA RVVAIISVFV IL LSIVIFC LETLPEFKHY KVFNTTTNGT KIEEDEVPDI TDPFFLIETL CIIWFTFELT VRFLACPNKL NFCRDVMNVI DII AIIPYF ITLATVVAEE EDTLNLPKAP VSPQDKSSNQ AMSLAILRVI RLVRVFRIFK LSRHSKGLQI LGRTLKASMR ELGL LIFFL FIGVVLFSSA VYFAEAGSEN SFFKSIPDAF WWAVVTMTTV GYGDMTPVGV WGKIVGSLCA IAGVLTIALP VPVIV SNFN YFYHRETDQE EMQSQNFNHV TSCPYLPGTL VGQHMKKSSL SESSSDMMDL DDGVESTPGL TETHPGRSAV APFLGA QQQ QQQPVASSLS MSIDKQLQHP LQQLTQTQLY QQQQQQQQQQ QNGFKQQQQQ TQQQLQQQQS HTINASAAAA TSGSGSS GL TMRHNNALAV SIETDV UniProtKB: Potassium voltage-gated channel protein Shaker |
-Macromolecule #2: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 2 / Number of copies: 32 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Macromolecule #3: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 9 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 289 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.39 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 663447 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X