登録情報 データベース : EMDB / ID : EMD-40707タイトル SARS-CoV-2 replication-transcription complex bound to RNA-nsp9, as a noncatalytic RNA-nsp9 binding mode Composite sharpened map 複合体 : SARS-CoV-2 replication-transcription complex bound to RNA-nsp9, as a noncatalytic RNA-nsp9 binding modeタンパク質・ペプチド : x 4種RNA : x 3種リガンド : x 4種 / / / / / / / / 機能・相同性 分子機能 ドメイン・相同性 構成要素
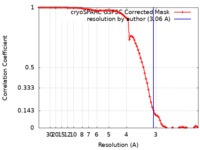
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 / 手法 / / 解像度 : 3.06 Å Small GI / Darst SA / Campbell EA 資金援助 Organization Grant number 国 National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID)
ジャーナル : Mol Cell / 年 : 2023タイトル : Structural and functional insights into the enzymatic plasticity of the SARS-CoV-2 NiRAN domain.著者 : Gabriel I Small / Olga Fedorova / Paul Dominic B Olinares / Joshua Chandanani / Anoosha Banerjee / Young Joo Choi / Henrik Molina / Brian T Chait / Seth A Darst / Elizabeth A Campbell / 要旨 : The enzymatic activity of the SARS-CoV-2 nidovirus RdRp-associated nucleotidyltransferase (NiRAN) domain is essential for viral propagation, with three distinct activities associated with ... The enzymatic activity of the SARS-CoV-2 nidovirus RdRp-associated nucleotidyltransferase (NiRAN) domain is essential for viral propagation, with three distinct activities associated with modification of the nsp9 N terminus, NMPylation, RNAylation, and deRNAylation/capping via a GDP-polyribonucleotidyltransferase reaction. The latter two activities comprise an unconventional mechanism for initiating viral RNA 5' cap formation, while the role of NMPylation is unclear. The structural mechanisms for these diverse enzymatic activities have not been properly delineated. Here, we determine high-resolution cryoelectron microscopy (cryo-EM) structures of catalytic intermediates for the NMPylation and deRNAylation/capping reactions, revealing diverse nucleotide binding poses and divalent metal ion coordination sites to promote its repertoire of activities. The deRNAylation/capping structure explains why GDP is a preferred substrate for the capping reaction over GTP. Altogether, these findings enhance our understanding of the promiscuous coronaviral NiRAN domain, a therapeutic target, and provide an accurate structural platform for drug development. 履歴 登録 2023年5月4日 - ヘッダ(付随情報) 公開 2023年11月22日 - マップ公開 2023年11月22日 - 更新 2023年11月22日 - 現状 2023年11月22日 処理サイト : RCSB / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報
 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報
 Severe acute respiratory syndrome coronavirus (SARS コロナウイルス)
Severe acute respiratory syndrome coronavirus (SARS コロナウイルス) データ登録者
データ登録者 米国, 1件
米国, 1件  引用
引用 ジャーナル: Mol Cell / 年: 2023
ジャーナル: Mol Cell / 年: 2023
 構造の表示
構造の表示 ダウンロードとリンク
ダウンロードとリンク emd_40707.map.gz
emd_40707.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-40707-v30.xml
emd-40707-v30.xml emd-40707.xml
emd-40707.xml EMDBヘッダ
EMDBヘッダ emd_40707_fsc.xml
emd_40707_fsc.xml FSCデータファイル
FSCデータファイル emd_40707.png
emd_40707.png emd-40707.cif.gz
emd-40707.cif.gz emd_40707_additional_1.map.gz
emd_40707_additional_1.map.gz emd_40707_additional_2.map.gz
emd_40707_additional_2.map.gz emd_40707_additional_3.map.gz
emd_40707_additional_3.map.gz emd_40707_half_map_1.map.gz
emd_40707_half_map_1.map.gz emd_40707_half_map_2.map.gz
emd_40707_half_map_2.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-40707
http://ftp.pdbj.org/pub/emdb/structures/EMD-40707 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40707
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40707 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_40707.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_40707.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 ムービー
ムービー コントローラー
コントローラー


















 Z
Z Y
Y X
X