[English] 日本語
 Yorodumi
Yorodumi- EMDB-4070: Structure-function insights reveal the human ribosome as a cancer... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4070 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure-function insights reveal the human ribosome as a cancer target for antibiotics | |||||||||
 Map data Map data | Chimera contour level | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CryoEM / human ribosome / 80S / cycloheximide / ribosome | |||||||||
| Function / homology |  Function and homology information Function and homology informationtranslation at presynapse / exit from mitosis / optic nerve development / response to insecticide / regulation of translation involved in cellular response to UV / eukaryotic 80S initiation complex / negative regulation of endoplasmic reticulum unfolded protein response / ribosomal protein import into nucleus / negative regulation of formation of translation preinitiation complex / axial mesoderm development ...translation at presynapse / exit from mitosis / optic nerve development / response to insecticide / regulation of translation involved in cellular response to UV / eukaryotic 80S initiation complex / negative regulation of endoplasmic reticulum unfolded protein response / ribosomal protein import into nucleus / negative regulation of formation of translation preinitiation complex / axial mesoderm development / regulation of G1 to G0 transition / retinal ganglion cell axon guidance / oxidized pyrimidine DNA binding / response to TNF agonist / positive regulation of base-excision repair / positive regulation of ubiquitin-protein transferase activity / positive regulation of respiratory burst involved in inflammatory response / protein-DNA complex disassembly / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / positive regulation of gastrulation / protein tyrosine kinase inhibitor activity / positive regulation of DNA-templated transcription initiation / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage / 90S preribosome assembly / IRE1-RACK1-PP2A complex / positive regulation of Golgi to plasma membrane protein transport / nucleolus organization / TNFR1-mediated ceramide production / alpha-beta T cell differentiation / positive regulation of DNA damage response, signal transduction by p53 class mediator / GAIT complex / negative regulation of RNA splicing / TORC2 complex binding / neural crest cell differentiation / supercoiled DNA binding / cytoplasmic translational initiation / NF-kappaB complex / negative regulation of DNA repair / G1 to G0 transition / oxidized purine DNA binding / cysteine-type endopeptidase activator activity involved in apoptotic process / middle ear morphogenesis / rRNA modification in the nucleus and cytosol / negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide / negative regulation of bicellular tight junction assembly / ubiquitin-like protein conjugating enzyme binding / regulation of establishment of cell polarity / negative regulation of phagocytosis / erythrocyte homeostasis / cytoplasmic side of rough endoplasmic reticulum membrane / Formation of the ternary complex, and subsequently, the 43S complex / ion channel inhibitor activity / protein kinase A binding / laminin receptor activity / Ribosomal scanning and start codon recognition / pigmentation / homeostatic process / positive regulation of mitochondrial depolarization / Translation initiation complex formation / macrophage chemotaxis / lung morphogenesis / negative regulation of Wnt signaling pathway / positive regulation of natural killer cell proliferation / male meiosis I / fibroblast growth factor binding / Protein hydroxylation / monocyte chemotaxis / BH3 domain binding / negative regulation of translational frameshifting / regulation of adenylate cyclase-activating G protein-coupled receptor signaling pathway / SARS-CoV-1 modulates host translation machinery / positive regulation of GTPase activity / TOR signaling / mTORC1-mediated signalling / iron-sulfur cluster binding / Peptide chain elongation / regulation of cell division / cellular response to ethanol / Selenocysteine synthesis / Formation of a pool of free 40S subunits / negative regulation of protein binding / positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator / Eukaryotic Translation Termination / protein serine/threonine kinase inhibitor activity / blastocyst development / SRP-dependent cotranslational protein targeting to membrane / Response of EIF2AK4 (GCN2) to amino acid deficiency / ubiquitin ligase inhibitor activity / negative regulation of respiratory burst involved in inflammatory response / Viral mRNA Translation / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / positive regulation of signal transduction by p53 class mediator / negative regulation of ubiquitin-dependent protein catabolic process / protein localization to nucleus / GTP hydrolysis and joining of the 60S ribosomal subunit / L13a-mediated translational silencing of Ceruloplasmin expression / Major pathway of rRNA processing in the nucleolus and cytosol / regulation of translational fidelity / protein targeting Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Myasnikov AG / Natchiar SK | |||||||||
 Citation Citation |  Journal: Nature / Year: 2015 Journal: Nature / Year: 2015Title: Structure of the human 80S ribosome. Authors: Heena Khatter / Alexander G Myasnikov / S Kundhavai Natchiar / Bruno P Klaholz /  Abstract: Ribosomes are translational machineries that catalyse protein synthesis. Ribosome structures from various species are known at the atomic level, but obtaining the structure of the human ribosome has ...Ribosomes are translational machineries that catalyse protein synthesis. Ribosome structures from various species are known at the atomic level, but obtaining the structure of the human ribosome has remained a challenge; efforts to address this would be highly relevant with regard to human diseases. Here we report the near-atomic structure of the human ribosome derived from high-resolution single-particle cryo-electron microscopy and atomic model building. The structure has an average resolution of 3.6 Å, reaching 2.9 Å resolution in the most stable regions. It provides unprecedented insights into ribosomal RNA entities and amino acid side chains, notably of the transfer RNA binding sites and specific molecular interactions with the exit site tRNA. It reveals atomic details of the subunit interface, which is seen to remodel strongly upon rotational movements of the ribosomal subunits. Furthermore, the structure paves the way for analysing antibiotic side effects and diseases associated with deregulated protein synthesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4070.map.gz emd_4070.map.gz | 23.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4070-v30.xml emd-4070-v30.xml emd-4070.xml emd-4070.xml | 109.4 KB 109.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4070.png emd_4070.png | 140.7 KB | ||
| Filedesc metadata |  emd-4070.cif.gz emd-4070.cif.gz | 21.4 KB | ||
| Others |  emd_4070_additional.map.gz emd_4070_additional.map.gz | 23.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4070 http://ftp.pdbj.org/pub/emdb/structures/EMD-4070 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4070 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4070 | HTTPS FTP |
-Related structure data
| Related structure data |  5lksMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4070.map.gz / Format: CCP4 / Size: 371.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4070.map.gz / Format: CCP4 / Size: 371.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Chimera contour level | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
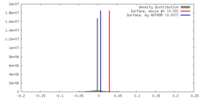
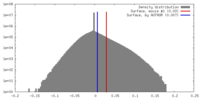
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Chimera contour level
| File | emd_4070_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Chimera contour level | ||||||||||||
| Projections & Slices |
| ||||||||||||
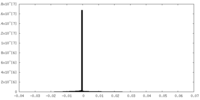
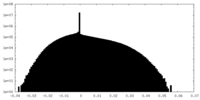
| Density Histograms |
- Sample components
Sample components
+Entire : Human 80S ribsome
+Supramolecule #1: Human 80S ribsome
+Macromolecule #1: 28S ribosomal RNA
+Macromolecule #2: 5S ribosomal RNA
+Macromolecule #3: 5.8S ribosomal RNA
+Macromolecule #47: 18S ribosomal RNA
+Macromolecule #4: 60S ribosomal protein L8
+Macromolecule #5: 60S ribosomal protein L3
+Macromolecule #6: 60S ribosomal protein L4
+Macromolecule #7: 60S ribosomal protein L5
+Macromolecule #8: 60S ribosomal protein L6
+Macromolecule #9: 60S ribosomal protein L7
+Macromolecule #10: 60S ribosomal protein L7a
+Macromolecule #11: 60S ribosomal protein L9
+Macromolecule #12: 60S ribosomal protein L10-like
+Macromolecule #13: 60S ribosomal protein L11
+Macromolecule #14: 60S ribosomal protein L13
+Macromolecule #15: 60S ribosomal protein L14
+Macromolecule #16: 60S ribosomal protein L15
+Macromolecule #17: 60S ribosomal protein L13a
+Macromolecule #18: 60S ribosomal protein L17
+Macromolecule #19: 60S ribosomal protein L18
+Macromolecule #20: 60S ribosomal protein L19
+Macromolecule #21: 60S ribosomal protein L18a
+Macromolecule #22: 60S ribosomal protein L21
+Macromolecule #23: 60S ribosomal protein L22
+Macromolecule #24: 60S ribosomal protein L23
+Macromolecule #25: 60S ribosomal protein L24
+Macromolecule #26: 60S ribosomal protein L23a
+Macromolecule #27: 60S ribosomal protein L26
+Macromolecule #28: 60S ribosomal protein L27
+Macromolecule #29: 60S ribosomal protein L27a
+Macromolecule #30: Ribosomal protein L29, isoform CRA_a
+Macromolecule #31: 60S ribosomal protein L30
+Macromolecule #32: 60S ribosomal protein L31
+Macromolecule #33: 60S ribosomal protein L32
+Macromolecule #34: 60S ribosomal protein L35a
+Macromolecule #35: 60S ribosomal protein L34
+Macromolecule #36: 60S ribosomal protein L35
+Macromolecule #37: 60S ribosomal protein L36
+Macromolecule #38: 60S ribosomal protein L37
+Macromolecule #39: 60S ribosomal protein L38
+Macromolecule #40: 60S ribosomal protein L39
+Macromolecule #41: Ubiquitin-60S ribosomal protein L40
+Macromolecule #42: 60S ribosomal protein L41
+Macromolecule #43: 60S ribosomal protein L36a
+Macromolecule #44: 60S ribosomal protein L37a
+Macromolecule #45: 60S ribosomal protein L28
+Macromolecule #46: 60S ribosomal protein L10a
+Macromolecule #48: 40S ribosomal protein SA
+Macromolecule #49: 40S ribosomal protein S3a
+Macromolecule #50: 40S ribosomal protein S3
+Macromolecule #51: 40S ribosomal protein S4, X isoform
+Macromolecule #52: 40S ribosomal protein S5
+Macromolecule #53: 40S ribosomal protein S7
+Macromolecule #54: 40S ribosomal protein S8
+Macromolecule #55: 40S ribosomal protein S10
+Macromolecule #56: 40S ribosomal protein S11
+Macromolecule #57: 40S ribosomal protein S15
+Macromolecule #58: 40S ribosomal protein S16
+Macromolecule #59: 40S ribosomal protein S17
+Macromolecule #60: 40S ribosomal protein S18
+Macromolecule #61: 40S ribosomal protein S19
+Macromolecule #62: 40S ribosomal protein S20
+Macromolecule #63: 40S ribosomal protein S21
+Macromolecule #64: 40S ribosomal protein S23
+Macromolecule #65: 40S ribosomal protein S26
+Macromolecule #66: 40S ribosomal protein S28
+Macromolecule #67: 40S ribosomal protein S29
+Macromolecule #68: Receptor of activated protein C kinase 1
+Macromolecule #69: 40S ribosomal protein S2
+Macromolecule #70: 40S ribosomal protein S6
+Macromolecule #71: 40S ribosomal protein S9
+Macromolecule #72: 40S ribosomal protein S12
+Macromolecule #73: 40S ribosomal protein S13
+Macromolecule #74: 40S ribosomal protein S14
+Macromolecule #75: 40S ribosomal protein S15a
+Macromolecule #76: 40S ribosomal protein S24
+Macromolecule #77: 40S ribosomal protein S25
+Macromolecule #78: 40S ribosomal protein S27
+Macromolecule #79: Ribosomal protein S30
+Macromolecule #80: Ubiquitin-40S ribosomal protein S27a
+Macromolecule #81: 4-{(2R)-2-[(1S,3S,5S)-3,5-dimethyl-2-oxocyclohexyl]-2-hydroxyethy...
+Macromolecule #82: MAGNESIUM ION
+Macromolecule #83: ZINC ION
+Macromolecule #84: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER/RHODIUM / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 79.0 K / Max: 105.0 K |
| Specialist optics | Spherical aberration corrector: CS corrected microscope |
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: OTHER / Digitization - Frames/image: 3-9 / Number grids imaged: 4 / Number real images: 6600 / Average exposure time: 1.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 3.0 µm / Calibrated defocus min: 0.5 µm / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 0.001 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 79000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT / Target criteria: Maximum likelihood |
|---|---|
| Output model |  PDB-5lks: |
 Movie
Movie Controller
Controller














































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)