[English] 日本語
 Yorodumi
Yorodumi- EMDB-37758: Fe-O nanocluster of form-XI in the 4-fold channel of Ureaplasma d... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Fe-O nanocluster of form-XI in the 4-fold channel of Ureaplasma diversum ferritin | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ferritin / METAL BINDING PROTEIN | |||||||||
| Biological species |  Ureaplasma diversum (bacteria) Ureaplasma diversum (bacteria) | |||||||||
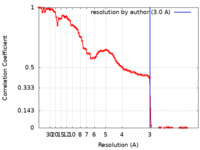
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Wang WM / Ma DY / Gong WJ / Wu LJ / Wang HF | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: J Am Chem Soc / Year: 2024 Journal: J Am Chem Soc / Year: 2024Title: Growth Process of Fe-O Nanoclusters with Different Sizes Biosynthesized by Protein Nanocages. Authors: Wenming Wang / Hongfang Xi / Dan Fu / Danyang Ma / Wenjun Gong / Yaqin Zhao / Xiaomei Li / Lijie Wu / Yu Guo / Guanghua Zhao / Hongfei Wang /  Abstract: All protein-directed syntheses of metal nanoclusters (NCs) and nanoparticles (NPs) have attracted considerable attention because protein scaffolds provide a unique metal coordination environment and ...All protein-directed syntheses of metal nanoclusters (NCs) and nanoparticles (NPs) have attracted considerable attention because protein scaffolds provide a unique metal coordination environment and can adjust the shape and morphology of NCs and NPs. However, the detailed formation mechanisms of NCs or NPs directed by protein templates remain unclear. In this study, by taking advantage of the ferritin nanocage as a biotemplate to monitor the growth of Fe-O NCs as a function of time, we synthesized a series of iron NCs with different sizes and shapes and subsequently solved their corresponding three-dimensional atomic-scale structures by X-ray protein crystallography and cryo-electron microscopy. The time-dependent structure analyses revealed the growth process of these Fe-O NCs with the 4-fold channel of ferritin as nucleation sites. To our knowledge, the newly biosynthesized FeOGlu represents the largest Fe-O NCs with a definite atomic structure. This study contributes to our understanding of the formation mechanism of iron NCs and provides an effective method for metal NC synthesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37758.map.gz emd_37758.map.gz | 59.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37758-v30.xml emd-37758-v30.xml emd-37758.xml emd-37758.xml | 14.3 KB 14.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37758_fsc.xml emd_37758_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_37758.png emd_37758.png | 144.9 KB | ||
| Filedesc metadata |  emd-37758.cif.gz emd-37758.cif.gz | 5.3 KB | ||
| Others |  emd_37758_half_map_1.map.gz emd_37758_half_map_1.map.gz emd_37758_half_map_2.map.gz emd_37758_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37758 http://ftp.pdbj.org/pub/emdb/structures/EMD-37758 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37758 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37758 | HTTPS FTP |
-Validation report
| Summary document |  emd_37758_validation.pdf.gz emd_37758_validation.pdf.gz | 1.3 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_37758_full_validation.pdf.gz emd_37758_full_validation.pdf.gz | 1.3 MB | Display | |
| Data in XML |  emd_37758_validation.xml.gz emd_37758_validation.xml.gz | 16.4 KB | Display | |
| Data in CIF |  emd_37758_validation.cif.gz emd_37758_validation.cif.gz | 21.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37758 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37758 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37758 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37758 | HTTPS FTP |
-Related structure data
| Related structure data |  8wqyMC  8w6mC  8w6qC  8w6sC  8w6uC  8w6yC  8w73C  8w74C  8w79C  8w7bC  8w7oC  8w7qC  8w7tC  8w7uC  8w7vC  8wptC  8wpvC  8wquC  8wqvC  8wqxC  8wr0C M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37758.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37758.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0469 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_37758_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_37758_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ferritin
| Entire | Name: ferritin |
|---|---|
| Components |
|
-Supramolecule #1: ferritin
| Supramolecule | Name: ferritin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Ureaplasma diversum (bacteria) Ureaplasma diversum (bacteria) |
-Macromolecule #1: ferritin
| Macromolecule | Name: ferritin / type: protein_or_peptide / ID: 1 / Details: WP_081847832.1 / Number of copies: 24 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Ureaplasma diversum (bacteria) Ureaplasma diversum (bacteria) |
| Molecular weight | Theoretical: 21.254996 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLYRERNNIM QKSNKINDAL NQHYKLNVEL GLVYAHYAHV ADDEFDMPYL GKFIQHLSED KLGVHKEYIS DYFKRNGMKL KTDVSVAVK SIPSDAKALI QEVYARENEV RDHVKAIAKL ALAEDDYESF YFIQWYVRDG LKDLTEVDDV VKLFNSSNDK L IIEETIKE MVEKEESEHE IWG |
-Macromolecule #2: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 2 / Number of copies: 24 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 10 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)