+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3773 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | High resolution INO80 core complex | |||||||||
 Map data Map data | High resolution human INO80 core complex. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Chromatin remodelling complex / GENE REGULATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationpromoter-enhancer loop anchoring activity / telomerase RNA localization to Cajal body / regulation of DNA strand elongation / positive regulation of telomere maintenance in response to DNA damage / establishment of protein localization to chromatin / R2TP complex / dynein axonemal particle / RPAP3/R2TP/prefoldin-like complex / Swr1 complex / Ino80 complex ...promoter-enhancer loop anchoring activity / telomerase RNA localization to Cajal body / regulation of DNA strand elongation / positive regulation of telomere maintenance in response to DNA damage / establishment of protein localization to chromatin / R2TP complex / dynein axonemal particle / RPAP3/R2TP/prefoldin-like complex / Swr1 complex / Ino80 complex / regulation of double-strand break repair / box C/D snoRNP assembly / protein folding chaperone complex / regulation of chromosome organization / NuA4 histone acetyltransferase complex / TFIID-class transcription factor complex binding / regulation of DNA replication / MLL1 complex / regulation of embryonic development / Telomere Extension By Telomerase / RNA polymerase II core promoter sequence-specific DNA binding / regulation of DNA repair / positive regulation of double-strand break repair via homologous recombination / Deposition of new CENPA-containing nucleosomes at the centromere / DNA helicase activity / telomere maintenance / TBP-class protein binding / positive regulation of DNA repair / cellular response to estradiol stimulus / euchromatin / negative regulation of canonical Wnt signaling pathway / Formation of the beta-catenin:TCF transactivating complex / ADP binding / beta-catenin binding / chromatin DNA binding / DNA Damage Recognition in GG-NER / nuclear matrix / cellular response to UV / transcription corepressor activity / UCH proteinases / unfolded protein binding / positive regulation of canonical Wnt signaling pathway / protein folding / nucleosome / HATs acetylate histones / ATPase binding / regulation of apoptotic process / DNA recombination / spermatogenesis / DNA helicase / transcription coactivator activity / regulation of cell cycle / Ub-specific processing proteases / protein stabilization / nuclear speck / ciliary basal body / cadherin binding / RNA polymerase II cis-regulatory region sequence-specific DNA binding / chromatin remodeling / ribonucleoprotein complex / cell division / DNA repair / centrosome / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / positive regulation of DNA-templated transcription / protein homodimerization activity / positive regulation of transcription by RNA polymerase II / ATP hydrolysis activity / extracellular exosome / nucleoplasm / ATP binding / identical protein binding / nucleus / membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.06 Å | |||||||||
 Authors Authors | Aramayo RJ / Bythell-Douglas R / Ayala R / Willhoft O / Wigley D / Zhang X | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2018 Journal: Nat Struct Mol Biol / Year: 2018Title: Cryo-EM structures of the human INO80 chromatin-remodeling complex. Authors: Ricardo J Aramayo / Oliver Willhoft / Rafael Ayala / Rohan Bythell-Douglas / Dale B Wigley / Xiaodong Zhang /  Abstract: Access to chromatin for processes such as transcription and DNA repair requires the sliding of nucleosomes along DNA. This process is aided by chromatin-remodeling complexes, such as the multisubunit ...Access to chromatin for processes such as transcription and DNA repair requires the sliding of nucleosomes along DNA. This process is aided by chromatin-remodeling complexes, such as the multisubunit INO80 chromatin-remodeling complex. Here we present cryo-EM structures of the active core complex of human INO80 at 9.6 Å, with portions at 4.1-Å resolution, and reconstructions of combinations of subunits. Together, these structures reveal the architecture of the INO80 complex, including Ino80 and actin-related proteins, which is assembled around a single RUVBL1 (Tip49a) and RUVBL2 (Tip49b) AAA+ heterohexamer. An unusual spoked-wheel structural domain of the Ino80 subunit is engulfed by this heterohexamer; both, in combination, form the core of the complex. We also identify a cleft in RUVBL1 and RUVBL2, which forms a major interaction site for partner proteins and probably communicates these interactions to its nucleotide-binding sites. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3773.map.gz emd_3773.map.gz | 9.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3773-v30.xml emd-3773-v30.xml emd-3773.xml emd-3773.xml | 13.2 KB 13.2 KB | Display Display |  EMDB header EMDB header |
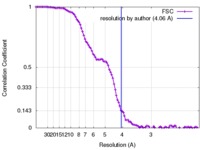
| FSC (resolution estimation) |  emd_3773_fsc.xml emd_3773_fsc.xml | 11.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_3773.png emd_3773.png | 51.4 KB | ||
| Filedesc metadata |  emd-3773.cif.gz emd-3773.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3773 http://ftp.pdbj.org/pub/emdb/structures/EMD-3773 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3773 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3773 | HTTPS FTP |
-Related structure data
| Related structure data |  5oafMC  3772C  3774C  3775C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3773.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3773.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | High resolution human INO80 core complex. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human INO80 core complex
| Entire | Name: Human INO80 core complex |
|---|---|
| Components |
|
-Supramolecule #1: Human INO80 core complex
| Supramolecule | Name: Human INO80 core complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 800 KDa |
-Macromolecule #1: RuvB-like 1
| Macromolecule | Name: RuvB-like 1 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO / EC number: DNA helicase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50.296914 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKIEEVKSTT KTQRIASHSH VKGLGLDESG LAKQAASGLV GQENAREACG VIVELIKSKK MAGRAVLLAG PPGTGKTALA LAIAQELGS KVPFCPMVGS EVYSTEIKKT EVLMENFRRA IGLRIKETKE VYEGEVTELT PCETENPMGG YGKTISHVII G LKTAKGTK ...String: MKIEEVKSTT KTQRIASHSH VKGLGLDESG LAKQAASGLV GQENAREACG VIVELIKSKK MAGRAVLLAG PPGTGKTALA LAIAQELGS KVPFCPMVGS EVYSTEIKKT EVLMENFRRA IGLRIKETKE VYEGEVTELT PCETENPMGG YGKTISHVII G LKTAKGTK QLKLDPSIFE SLQKERVEAG DVIYIEANSG AVKRQGRCDT YATEFDLEAE EYVPLPKGDV HKKKEIIQDV TL HDLDVAN ARPQGGQDIL SMMGQLMKPK KTEITDKLRG EINKVVNKYI DQGIAELVPG VLFVDEVHML DIECFTYLHR ALE SSIAPI VIFASNRGNC VIRGTEDITS PHGIPLDLLD RVMIIRTMLY TPQEMKQIIK IRAQTEGINI SEEALNHLGE IGTK TTLRY SVQLLTPANL LAKINGKDSI EKEHVEEISE LFYDAKSSAK ILADQQDKYM K UniProtKB: RuvB-like 1 |
-Macromolecule #2: RuvB-like 2
| Macromolecule | Name: RuvB-like 2 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO / EC number: DNA helicase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 51.222465 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MATVTATTKV PEIRDVTRIE RIGAHSHIRG LGLDDALEPR QASQGMVGQL AARRAAGVVL EMIREGKIAG RAVLIAGQPG TGKTAIAMG MAQALGPDTP FTAIAGSEIF SLEMSKTEAL TQAFRRSIGV RIKEETEIIE GEVVEIQIDR PATGTGSKVG K LTLKTTEM ...String: MATVTATTKV PEIRDVTRIE RIGAHSHIRG LGLDDALEPR QASQGMVGQL AARRAAGVVL EMIREGKIAG RAVLIAGQPG TGKTAIAMG MAQALGPDTP FTAIAGSEIF SLEMSKTEAL TQAFRRSIGV RIKEETEIIE GEVVEIQIDR PATGTGSKVG K LTLKTTEM ETIYDLGTKM IESLTKDKVQ AGDVITIDKA TGKISKLGRS FTRARDYDAM GSQTKFVQCP DGELQKRKEV VH TVSLHEI DVINSRTQGF LALFSGDTGE IKSEVREQIN AKVAEWREEG KAEIIPGVLF IDEVHMLDIE SFSFLNRALE SDM APVLIM ATNRGITRIR GTSYQSPHGI PIDLLDRLLI VSTTPYSEKD TKQILRIRCE EEDVEMSEDA YTVLTRIGLE TSLR YAIQL ITAASLVCRK RKGTEVQVDD IKRVYSLFLD ESRSTQYMKE YQDAFLFNEL KGETMDTS UniProtKB: RuvB-like 2 |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 6 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-5oaf: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)