[English] 日本語
 Yorodumi
Yorodumi- EMDB-37130: Cryo-EM structure of the human parainfluenza virus hPIV3 L-P poly... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in dimeric form | |||||||||
 Map data Map data | Cryo-EM map of the human parainfluenza virus hPIV3 L-P polymerase in dimeric form, class 1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | dimeric polymerase / parainfluenza virus / L-P polymerase / cryo-EM structure / non-segmented negative-strand RNA virus / L-L dimerization / RNA replication / RdRp active site / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationGDP polyribonucleotidyltransferase / Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides / viral genome replication / Transferases; Transferring one-carbon groups; Methyltransferases / virion component / mRNA 5'-cap (guanine-N7-)-methyltransferase activity / host cell cytoplasm / RNA-directed RNA polymerase / RNA-dependent RNA polymerase activity / GTPase activity ...GDP polyribonucleotidyltransferase / Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides / viral genome replication / Transferases; Transferring one-carbon groups; Methyltransferases / virion component / mRNA 5'-cap (guanine-N7-)-methyltransferase activity / host cell cytoplasm / RNA-directed RNA polymerase / RNA-dependent RNA polymerase activity / GTPase activity / DNA-templated transcription / RNA binding / ATP binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Human respirovirus 3 Human respirovirus 3 | |||||||||
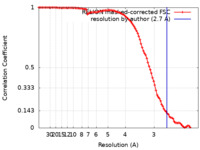
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | |||||||||
 Authors Authors | Xie J / Wang L / Zhai G / Wu D / Lin Z / Wang M / Yan X / Gao L / Huang X / Fearns R / Chen S | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural basis for dimerization of a paramyxovirus polymerase complex. Authors: Jin Xie / Mohamed Ouizougun-Oubari / Li Wang / Guanglei Zhai / Daitze Wu / Zhaohu Lin / Manfu Wang / Barbara Ludeke / Xiaodong Yan / Tobias Nilsson / Lu Gao / Xinyi Huang / Rachel Fearns / Shuai Chen /    Abstract: The transcription and replication processes of non-segmented, negative-strand RNA viruses (nsNSVs) are catalyzed by a multi-functional polymerase complex composed of the large protein (L) and a ...The transcription and replication processes of non-segmented, negative-strand RNA viruses (nsNSVs) are catalyzed by a multi-functional polymerase complex composed of the large protein (L) and a cofactor protein, such as phosphoprotein (P). Previous studies have shown that the nsNSV polymerase can adopt a dimeric form, however, the structure of the dimer and its function are poorly understood. Here we determine a 2.7 Å cryo-EM structure of human parainfluenza virus type 3 (hPIV3) L-P complex with the connector domain (CD') of a second L built, while reconstruction of the rest of the second L-P obtains a low-resolution map of the ring-like L core region. This study reveals detailed atomic features of nsNSV polymerase active site and distinct conformation of hPIV3 L with a unique β-strand latch. Furthermore, we report the structural basis of L-L dimerization, with CD' located at the putative template entry of the adjoining L. Disruption of the L-L interface causes a defect in RNA replication that can be overcome by complementation, demonstrating that L dimerization is necessary for hPIV3 genome replication. These findings provide further insight into how nsNSV polymerases perform their functions, and suggest a new avenue for rational drug design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37130.map.gz emd_37130.map.gz | 42.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37130-v30.xml emd-37130-v30.xml emd-37130.xml emd-37130.xml | 19.5 KB 19.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37130_fsc.xml emd_37130_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_37130.png emd_37130.png | 43.4 KB | ||
| Masks |  emd_37130_msk_1.map emd_37130_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-37130.cif.gz emd-37130.cif.gz | 7.5 KB | ||
| Others |  emd_37130_half_map_1.map.gz emd_37130_half_map_1.map.gz emd_37130_half_map_2.map.gz emd_37130_half_map_2.map.gz | 77.3 MB 77.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37130 http://ftp.pdbj.org/pub/emdb/structures/EMD-37130 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37130 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37130 | HTTPS FTP |
-Validation report
| Summary document |  emd_37130_validation.pdf.gz emd_37130_validation.pdf.gz | 791.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_37130_full_validation.pdf.gz emd_37130_full_validation.pdf.gz | 791.5 KB | Display | |
| Data in XML |  emd_37130_validation.xml.gz emd_37130_validation.xml.gz | 19.5 KB | Display | |
| Data in CIF |  emd_37130_validation.cif.gz emd_37130_validation.cif.gz | 25.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37130 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37130 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37130 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37130 | HTTPS FTP |
-Related structure data
| Related structure data |  8kdbMC  8kdcC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37130.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37130.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of the human parainfluenza virus hPIV3 L-P polymerase in dimeric form, class 1 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.087 Å | ||||||||||||||||||||||||||||||||||||
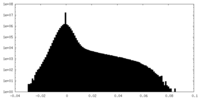
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_37130_msk_1.map emd_37130_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
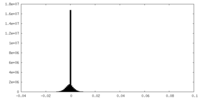
| Density Histograms |
-Half map: half1 map
| File | emd_37130_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half1 map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half2 map
| File | emd_37130_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half2 map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human parainfluenza virus hPIV3 L-P polymerase in dimeric form
| Entire | Name: Human parainfluenza virus hPIV3 L-P polymerase in dimeric form |
|---|---|
| Components |
|
-Supramolecule #1: Human parainfluenza virus hPIV3 L-P polymerase in dimeric form
| Supramolecule | Name: Human parainfluenza virus hPIV3 L-P polymerase in dimeric form type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Human respirovirus 3 Human respirovirus 3 |
-Macromolecule #1: RNA-directed RNA polymerase L
| Macromolecule | Name: RNA-directed RNA polymerase L / type: protein_or_peptide / ID: 1 / Details: NP_067153.2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human respirovirus 3 Human respirovirus 3 |
| Molecular weight | Theoretical: 260.103703 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MISNQQSDNG QKENIKNLGA KRARKMDTES NNGTVSDILY PECHLNSPIV KGKIAQLHTI MSLPQPYDMD DDSILVITRQ KIKLNKLDK RQRSIRRLKL ILTEKVNDLG KYTFIRYPEM SKEMFKLYIP GINSKVTELL LKADRTYSQM TDGLRDLWIN V LSKLASKN ...String: MISNQQSDNG QKENIKNLGA KRARKMDTES NNGTVSDILY PECHLNSPIV KGKIAQLHTI MSLPQPYDMD DDSILVITRQ KIKLNKLDK RQRSIRRLKL ILTEKVNDLG KYTFIRYPEM SKEMFKLYIP GINSKVTELL LKADRTYSQM TDGLRDLWIN V LSKLASKN DGSNYDLNEE INNISKVHTT YKSDKWYNPF KTWFTIKYDM RRLQKARNEI TFNVGKDYNL LEDQKNFLLI HP ELVLILD KQNYNGYLIT PELVLMYCDV VEGRWNISAC AKLDPKLQSM YQKGNNLWEV IDKLFPIMGE KTFDVISLLE PLA LSLIQT HDPVKQLRGA FLNHVLSEME LIFESGESIR EFLSVDYIDK ILDIFNESTI DEIAEIFSFF RTFGHPPLEA SIAA EKVRK YMYIEKQLKF DTVNKCHAIF CTIIINGYRE RHGGQWPPVT LPDHAHEFII NAYGSNSAIS YENAVDYYQS FIGIK FNKF IEPQLDEDLT IYMKDKALSP KKSNWDTVYP ASNLLYRTNA SNESRRLVEV FIADSKFDPH QILDYVESGD WLDDPE FNI SYSLKEKEIK QEGRLFAKMT YKMRATQVLS ETLLANNIGK FFQENGMVKG EIELLKRLTT ISISGVPRYN EVYNNSK SH TDDLKTYNKI SNLNLSSNQK SKKFEFKSTD IYNDGYETVS CFLTTDLKKY CLNWRYESTA LFGETCNQIF GLNKLFNW L HPRLEGSTIY VGDPYCPPSD KEHISLEDHP DSGFYVHNPR GGIEGFCQKL WTLISISAIH LAAVRIGVRV TAMVQGDNQ AIAVTTRVPN NYDYRIKKEI VYKDVVRFFD SLREVMDDLG HELKLNETII SSKMFIYSKR IYYDGRILPQ ALKALSRCVF WSETVIDET RSASSNLATS FAKAIENGYS PVLGYACSIF KNIQQLYIAL GMNINPTITQ NIKDQYFKNS NWMQYASLIP A SVGGFNYM AMSRCFVRNI GDPSVAALAD IKRFIKANLL DRSVLYRIMN QEPGESSFLD WASDPYSCNL PQSQNITTMI KN ITARNVL QDSPNPLLSG LFTNTMIEED EELAEFLMDR KVILPRVAHD ILDNSLTGIR NAIAGMLDTT KSLIRVGINR GGL TYSLLR KISNYDLVQY ETLSRTLRLI VSDKIRYEDM CSVDLAIALR QKMWIHLSGG RMISGLETPD PLELLSGVVI TGSE HCKIC YSSDGTNPYT WMYLPGNIKI GSAETGVSSL RVPYFGSVTD ERSEAQLGYI KNLSKPAKAA IRIAMIYTWA FGNDE ISWM EASQIAQTRA NFTLDSLKIL TPVATSTNLS HRLKDTATQM KFSSTSLIRV SRFITMSNDN MSIKEANETK DTNLIY QQI MLTGLSVFEY LFRLKETTGH NPIVMHLHIE DECCIKESFN DEHINPESTL ELIRYPESNE FIYDKDPLKD VDLSKLM VI KDHSYTIDMN YWDDTDIIHA ISICTAITIA DTMSQLDRDN LKEIIVIAND DDINSLITEF LTLDILVFLK TFGGLLVN Q FAYTLYSLKI EGRDLIWDYI MRTLRDTSHS ILKVLSNALS HPKVFKRFWD CGVLNPIYGP NTASQDQIKL ALSICEYAL DLFMREWLNG VSLEIYICDS DMEVANDRKQ AFISRHLSFV CCLAEIASFG PNLLNLTYLE RLDLLKQYLE LNIKEDPTLK YVQISGLLI KSFPSTVTYV RKTAIKYLRI RGISPPEVID DWDPIEDENM LDNIVKTIND NCNKDNKGNK INNFWGLALK N YQVLKIRS ITSDSDDNDR LDASTSGLTL PQGGNYLSHQ LRLFGINSTS CLKALELSQI LMKEVNKDKD RLFLGEGAGA ML ACYDATL GPAINYYNSG LNITDVIGQR ELKIFPSEVS LVGKKLGNVT QILNRVKVLF NGNPNSTWIG NMECESLIWS ELN DKSIGL VHCDMEGAIG KSEETVLHEH YSVIRITYLI GDDDVVLVSK IIPTITPNWS RILYLYKLYW KDVSIISLKT SNPA STELY LISKDAYCTI MEPSEVVLSK LKRLSLLEEN NLLKWIILSK KRNNEWLHHE IKEGERDYGV MRPYHMALQI FGFQI NLNH LAKEFLSTPD LTNINNIIQS FQRTIKDVLF EWINITHDDK RHKLGGRYNI FPLKNKGKLR LLSRRLVLSW ISLSLS TRL LTGRFPDEKF EHRAQTGYVS LADTDLESLK LLSKNIIKNY RECIGSISYW FLTKEVKILM KLIGGAKLLG IPRQYKE PE EQLLENYNQH DEFDIDDYKD DDDK UniProtKB: RNA-directed RNA polymerase L |
-Macromolecule #2: Phosphoprotein
| Macromolecule | Name: Phosphoprotein / type: protein_or_peptide / ID: 2 / Details: NP_067149.1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human respirovirus 3 Human respirovirus 3 |
| Molecular weight | Theoretical: 68.471312 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MESDAKNYQI MDSWEEESRD KSTNISSALN IIEFILSTDP QEDLSENDTI NTRTQQLSAT IYQPKIKPTE TSEKDSGSTD KNRQSGSSH ECTTEAKDRT IDQETVQRGP GRRSSSDSRA ETVVSGGISR SITNSKNGTQ NTEDIDLNEI RKMDKDSIEG K VRQSADVP ...String: MESDAKNYQI MDSWEEESRD KSTNISSALN IIEFILSTDP QEDLSENDTI NTRTQQLSAT IYQPKIKPTE TSEKDSGSTD KNRQSGSSH ECTTEAKDRT IDQETVQRGP GRRSSSDSRA ETVVSGGISR SITNSKNGTQ NTEDIDLNEI RKMDKDSIEG K VRQSADVP SEISGSDVIF TTEQSRNSDH GRSLESISTP DTRSISVVTA ATPDDEEEIL MKNSRTKKSS SIHQEDDKRI KK GGKGKDW FKKSKDTDNQ IPTSDYRSTS KGQKKISKTT TINTDTKGQT EIQTESSGTQ SSSWNLTIDN NTDRTEQTNT TPP TTTSGS TYTKESIRTN SGSKPKTQKT NGKERKDTEE SNRFTERAIT LLQNLGVIQS TSKLDLYQDK RVVCVANVLN NVDT ASKID FLAGLVIGVS MDNDTKLTQI QNEMLNLKAD LKKMDESHRR LIENQREQLS LITSLISNLK IMTERGGKKD QNESN ERVS MIKTKLKEEK IKKTRFDPLM ETQGIDKNIP DLYRHAGNTL ENDVQVKSEI LSSYNESNAT RLIPKKVSST MRSLVA VIS NSNLSQSTKQ SYINELKHCK NDEEVSELMD MFNEDVNNCQ HHHHHH UniProtKB: Phosphoprotein |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #4: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)