[English] 日本語
 Yorodumi
Yorodumi- EMDB-35981: Single-particle cryo-EM structure of mouse apoferritin at 1.49 An... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
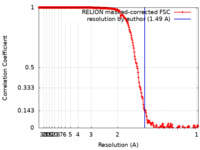
| Title | Single-particle cryo-EM structure of mouse apoferritin at 1.49 Angstrom resolution (Dataset B) | |||||||||
 Map data Map data | Postprocess_masked | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | single-particle cryo-EM / Cold field emission / CFEG / Apoferritin / CRYO ARM / STRUCTURAL PROTEIN | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 1.49 Å | |||||||||
 Authors Authors | Kawakami K / Maki-Yonekura S / Hamaguchi T / Takaba K / Yonekura K | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: Commun Chem / Year: 2023 Journal: Commun Chem / Year: 2023Title: Measurement of charges and chemical bonding in a cryo-EM structure. Authors: Saori Maki-Yonekura / Keisuke Kawakami / Kiyofumi Takaba / Tasuku Hamaguchi / Koji Yonekura /  Abstract: Hydrogen bonding, bond polarity, and charges in protein molecules play critical roles in the stabilization of protein structures, as well as affecting their functions such as enzymatic catalysis, ...Hydrogen bonding, bond polarity, and charges in protein molecules play critical roles in the stabilization of protein structures, as well as affecting their functions such as enzymatic catalysis, electron transfer, and ligand binding. These effects can potentially be measured in Coulomb potentials using cryogenic electron microscopy (cryo-EM). We here present charges and bond properties of hydrogen in a sub-1.2 Å resolution structure of a protein complex, apoferritin, by single-particle cryo-EM. A weighted difference map reveals positive densities for most hydrogen atoms in the core region of the complex, while negative densities around acidic amino-acid side chains are likely related to negative charges. The former positive densities identify the amino- and oxo-termini of asparagine and glutamine side chains. The latter observations were verified by spatial-resolution selection and a dose-dependent frame series. The average position of the hydrogen densities depends on the parent bonded-atom type, and this is validated by the estimated level of the standard uncertainties in the bond lengths. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35981.map.gz emd_35981.map.gz | 68.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35981-v30.xml emd-35981-v30.xml emd-35981.xml emd-35981.xml | 13.1 KB 13.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_35981_fsc.xml emd_35981_fsc.xml | 21.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_35981.png emd_35981.png | 111.6 KB | ||
| Others |  emd_35981_half_map_1.map.gz emd_35981_half_map_1.map.gz emd_35981_half_map_2.map.gz emd_35981_half_map_2.map.gz | 659 MB 659 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35981 http://ftp.pdbj.org/pub/emdb/structures/EMD-35981 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35981 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35981 | HTTPS FTP |
-Validation report
| Summary document |  emd_35981_validation.pdf.gz emd_35981_validation.pdf.gz | 635.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_35981_full_validation.pdf.gz emd_35981_full_validation.pdf.gz | 635.2 KB | Display | |
| Data in XML |  emd_35981_validation.xml.gz emd_35981_validation.xml.gz | 28.6 KB | Display | |
| Data in CIF |  emd_35981_validation.cif.gz emd_35981_validation.cif.gz | 38 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35981 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35981 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35981 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35981 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_35981.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35981.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocess_masked | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.495 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half1 map
| File | emd_35981_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half1_map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half2 map
| File | emd_35981_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half2_map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Apoferritin
| Entire | Name: Apoferritin |
|---|---|
| Components |
|
-Supramolecule #1: Apoferritin
| Supramolecule | Name: Apoferritin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Ferritin heavy chain
| Macromolecule | Name: Ferritin heavy chain / type: protein_or_peptide / ID: 1 / Enantiomer: DEXTRO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: PSQVRQNYHQ DAEAAINRQI NLELYASYVY LSMSCYFDRD DVALKNFAKY FLHQSHEERE HAEKLMKLQN QRGGRIFLQD IKKPDRDDWE SGLNAMECAL HLEKSVNQSL LELHKLATDK NDPHLCDFIE TYYLSEQVKS IKELGDHVTN LRKMGAPEAG MAEYLFDKHT LG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 51.82 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 150.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.0 µm / Nominal defocus min: 0.5 µm |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X