+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | S-ECD (Omicron XBB.1) in complex with PD of ACE2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-Cov-2 / VIRAL PROTEIN / VIRAL PROTEIN-HYDROLASE complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin / positive regulation of gap junction assembly / regulation of vasoconstriction / regulation of cardiac conduction ...positive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin / positive regulation of gap junction assembly / regulation of vasoconstriction / regulation of cardiac conduction / peptidyl-dipeptidase activity / maternal process involved in female pregnancy / angiotensin maturation / metallocarboxypeptidase activity / Metabolism of Angiotensinogen to Angiotensins / Attachment and Entry / carboxypeptidase activity / negative regulation of signaling receptor activity / positive regulation of cardiac muscle contraction / regulation of cytokine production / viral life cycle / blood vessel diameter maintenance / brush border membrane / regulation of transmembrane transporter activity / negative regulation of smooth muscle cell proliferation / negative regulation of ERK1 and ERK2 cascade / cilium / metallopeptidase activity / endocytic vesicle membrane / positive regulation of reactive oxygen species metabolic process / virus receptor activity / regulation of cell population proliferation / regulation of inflammatory response / Maturation of spike protein / viral translation / endopeptidase activity / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / Potential therapeutics for SARS / structural constituent of virion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / entry receptor-mediated virion attachment to host cell / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / membrane fusion / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / receptor ligand activity / host cell surface receptor binding / symbiont entry into host cell / membrane raft / apical plasma membrane / endoplasmic reticulum lumen / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / cell surface / extracellular space / zinc ion binding / extracellular exosome / extracellular region / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.95 Å | |||||||||
 Authors Authors | Li YN / Shen YP / Zhang YY / Yan RH | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural basis for the enhanced infectivity and immune evasion of Omicron subvariants Authors: Li Y / Shen YP / Zhang Y / Yan R | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35267.map.gz emd_35267.map.gz | 78.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35267-v30.xml emd-35267-v30.xml emd-35267.xml emd-35267.xml | 17 KB 17 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35267.png emd_35267.png | 31.1 KB | ||
| Filedesc metadata |  emd-35267.cif.gz emd-35267.cif.gz | 6.7 KB | ||
| Others |  emd_35267_half_map_1.map.gz emd_35267_half_map_1.map.gz emd_35267_half_map_2.map.gz emd_35267_half_map_2.map.gz | 84.7 MB 84.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35267 http://ftp.pdbj.org/pub/emdb/structures/EMD-35267 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35267 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35267 | HTTPS FTP |
-Validation report
| Summary document |  emd_35267_validation.pdf.gz emd_35267_validation.pdf.gz | 894.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_35267_full_validation.pdf.gz emd_35267_full_validation.pdf.gz | 894.2 KB | Display | |
| Data in XML |  emd_35267_validation.xml.gz emd_35267_validation.xml.gz | 13.2 KB | Display | |
| Data in CIF |  emd_35267_validation.cif.gz emd_35267_validation.cif.gz | 15.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35267 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35267 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35267 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35267 | HTTPS FTP |
-Related structure data
| Related structure data |  8i9dMC  8i9bC  8i9cC  8i9eC  8i9fC  8i9gC  8i9hC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35267.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35267.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.087 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35267_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
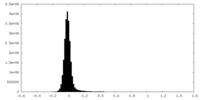
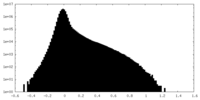
| Density Histograms |
-Half map: #1
| File | emd_35267_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
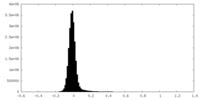
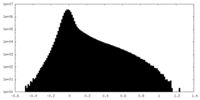
| Density Histograms |
- Sample components
Sample components
-Entire : S-ECD (Omicron XBB.1) in complex with PD of ACE2
| Entire | Name: S-ECD (Omicron XBB.1) in complex with PD of ACE2 |
|---|---|
| Components |
|
-Supramolecule #1: S-ECD (Omicron XBB.1) in complex with PD of ACE2
| Supramolecule | Name: S-ECD (Omicron XBB.1) in complex with PD of ACE2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #2: S-ECD (Omicron XBB.1)
| Supramolecule | Name: S-ECD (Omicron XBB.1) / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: PD of ACE2
| Supramolecule | Name: PD of ACE2 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Details: Omicron XBB.1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 140.591641 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPA LPFNDGVYF ASTEKSNIIR GWIFGTTLDS KTQSLLIVNN ATNVVIKVCE FQFCNDPFLD VYQKNNKSWM ESEFRVYSSA N NCTFEYVS ...String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPA LPFNDGVYF ASTEKSNIIR GWIFGTTLDS KTQSLLIVNN ATNVVIKVCE FQFCNDPFLD VYQKNNKSWM ESEFRVYSSA N NCTFEYVS QPFLMDLEGK EGNFKNLREF VFKNIDGYFK IYSKHTPINL ERDLPQGFSA LEPLVDLPIG INITRFQTLL AL HRSYLTP VDSSSGWTAG AAAYYVGYLQ PRTFLLKYNE NGTITDAVDC ALDPLSETKC TLKSFTVEKG IYQTSNFRVQ PTE SIVRFP NITNLCPFHE VFNATTFASV YAWNRKRISN CVADYSVIYN FAPFFAFKCY GVSPTKLNDL CFTNVYADSF VIRG NEVSQ IAPGQTGNIA DYNYKLPDDF TGCVIAWNSN KLDSKPSGNY NYLYRLFRKS KLKPFERDIS TEIYQAGNKP CNGVA GSNC YSPLQSYGFR PTYGVGHQPY RVVVLSFELL HAPATVCGPK KSTNLVKNKC VNFNFNGLTG TGVLTESNKK FLPFQQ FGR DIADTTDAVR DPQTLEILDI TPCSFGGVSV ITPGTNTSNQ VAVLYQGVNC TEVPVAIHAD QLTPTWRVYS TGSNVFQ TR AGCLIGAEYV NNSYECDIPI GAGICASYQT QTKSHGSASS VASQSIIAYT MSLGAENSVA YSNNSIAIPT NFTISVTT E ILPVSMTKTS VDCTMYICGD STECSNLLLQ YGSFCTQLKR ALTGIAVEQD KNTQEVFAQV KQIYKTPPIK YFGGFNFSQ ILPDPSKPSK RSPIEDLLFN KVTLADAGFI KQYGDCLGDI AARDLICAQK FNGLTVLPPL LTDEMIAQYT SALLAGTITS GWTFGAGPA LQIPFPMQMA YRFNGIGVTQ NVLYENQKLI ANQFNSAIGK IQDSLSSTPS ALGKLQDVVN HNAQALNTLV K QLSSKFGA ISSVLNDILS RLDPPEAEVQ IDRLITGRLQ SLQTYVTQQL IRAAEIRASA NLAATKMSEC VLGQSKRVDF CG KGYHLMS FPQSAPHGVV FLHVTYVPAQ EKNFTTAPAI CHDGKAHFPR EGVFVSNGTH WFVTQRNFYE PQIITTDNTF VSG NCDVVI GIVNNTVYDP LQPELDSFKE ELDKYFKNHT SPDVDLGDIS GINASVVNIQ KEIDRLNEVA KNLNESLIDL QELG KYEQG SGYIPEAPRD GQAYVRKDGE WVLLSTFLLE GSDEVDAGSH HHHHHHHHHG SVEDYKDDDD K UniProtKB: Spike glycoprotein |
-Macromolecule #2: Processed angiotensin-converting enzyme 2
| Macromolecule | Name: Processed angiotensin-converting enzyme 2 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 72.039992 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGVKVLFALI CIAVAEAGTS TIEEQAKTFL DKFNHEAEDL FYQSSLASWN YNTNITEENV QNMNNAGDKW SAFLKEQSTL AQMYPLQEI QNLTVKLQLQ ALQQNGSSVL SEDKSKRLNT ILNTMSTIYS TGKVCNPDNP QECLLLEPGL NEIMANSLDY N ERLWAWES ...String: MGVKVLFALI CIAVAEAGTS TIEEQAKTFL DKFNHEAEDL FYQSSLASWN YNTNITEENV QNMNNAGDKW SAFLKEQSTL AQMYPLQEI QNLTVKLQLQ ALQQNGSSVL SEDKSKRLNT ILNTMSTIYS TGKVCNPDNP QECLLLEPGL NEIMANSLDY N ERLWAWES WRSEVGKQLR PLYEEYVVLK NEMARANHYE DYGDYWRGDY EVNGVDGYDY SRGQLIEDVE HTFEEIKPLY EH LHAYVRA KLMNAYPSYI SPIGCLPAHL LGDMWGRFWT NLYSLTVPFG QKPNIDVTDA MVDQAWDAQR IFKEAEKFFV SVG LPNMTQ GFWENSMLTD PGNVQKAVCH PTAWDLGKGD FRILMCTKVT MDDFLTAHHE MGHIQYDMAY AAQPFLLRNG ANEG FHEAV GEIMSLSAAT PKHLKSIGLL SPDFQEDNET EINFLLKQAL TIVGTLPFTY MLEKWRWMVF KGEIPKDQWM KKWWE MKRE IVGVVEPVPH DETYCDPASL FHVSNDYSFI RYYTRTLYQF QFQEALCQAA KHEGPLHKCD ISNSTEAGQK LFNMLR LGK SEPWTLALEN VVGAKNMNVR PLLNYFEPLF TWLKDQNKNS FVGWSTDWSP YADDYKDDDD K UniProtKB: Angiotensin-converting enzyme 2 |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 28 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.95 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. v4) / Number images used: 804338 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller















 Z
Z Y
Y X
X