+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Echovirus3 full particle in complex with 6D10 Fab | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / cytoplasmic vesicle membrane / endocytosis involved in viral entry into host cell / viral capsid ...symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / cytoplasmic vesicle membrane / endocytosis involved in viral entry into host cell / viral capsid / nucleoside-triphosphate phosphatase / protein complex oligomerization / symbiont-mediated suppression of host gene expression / monoatomic ion channel activity / DNA replication / host cell cytoplasm / RNA helicase activity / symbiont entry into host cell / induction by virus of host autophagy / RNA-directed RNA polymerase / viral RNA genome replication / cysteine-type endopeptidase activity / RNA-dependent RNA polymerase activity / virus-mediated perturbation of host defense response / DNA-templated transcription / virion attachment to host cell / structural molecule activity / ATP hydrolysis activity / proteolysis / RNA binding / ATP binding / metal ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Echovirus E3 Echovirus E3 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Wang X / Fu W | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Viruses / Year: 2022 Journal: Viruses / Year: 2022Title: Structural Basis for the Immunogenicity of the C-Terminus of VP1 of Echovirus 3 Revealed by the Binding of a Neutralizing Antibody. Authors: Shuai Qi / Wangjun Fu / Jinyan Fan / Li Zhang / Binyang Zheng / Kang Wang / Xiangxi Wang / Ling Zhu / Xinjian Li / Yuxia Zhang /  Abstract: Echovirus 3 (E3), a serotype of human enterovirus B (HEV-B), causes severe diseases in infants. Here, we determined the structures of E3 with a monoclonal antibody (MAb) 6D10 by cryo-EM to ...Echovirus 3 (E3), a serotype of human enterovirus B (HEV-B), causes severe diseases in infants. Here, we determined the structures of E3 with a monoclonal antibody (MAb) 6D10 by cryo-EM to comprehensively understand the specificities and the immunological characteristic of this serotype. The solved cryo-EM structures of the F-, A-, and E-particles of E3 bound with 6D10 revealed the structural features of the virus-antibody interface. Importantly, the structures of E-particles bound with 6D10 revealed for the first time the nature of the C-terminus of VP1 for HEV-Bs at the structural level. The highly immunogenic nature of this region in the E-particles provides new strategies for vaccine development for HEV-Bs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34232.map.gz emd_34232.map.gz | 196.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34232-v30.xml emd-34232-v30.xml emd-34232.xml emd-34232.xml | 20.3 KB 20.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34232.png emd_34232.png | 61.2 KB | ||
| Others |  emd_34232_half_map_1.map.gz emd_34232_half_map_1.map.gz emd_34232_half_map_2.map.gz emd_34232_half_map_2.map.gz | 166.1 MB 166.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34232 http://ftp.pdbj.org/pub/emdb/structures/EMD-34232 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34232 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34232 | HTTPS FTP |
-Validation report
| Summary document |  emd_34232_validation.pdf.gz emd_34232_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_34232_full_validation.pdf.gz emd_34232_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_34232_validation.xml.gz emd_34232_validation.xml.gz | 16 KB | Display | |
| Data in CIF |  emd_34232_validation.cif.gz emd_34232_validation.cif.gz | 18.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34232 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34232 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34232 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34232 | HTTPS FTP |
-Related structure data
| Related structure data |  8gsdMC  8gscC  8gseC  8gsfC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34232.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34232.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.332 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34232_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
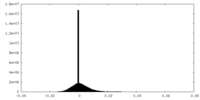
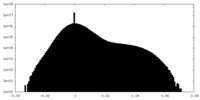
| Density Histograms |
-Half map: #1
| File | emd_34232_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
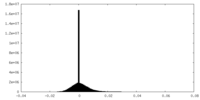
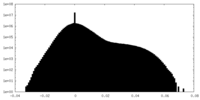
| Density Histograms |
- Sample components
Sample components
+Entire : Echovirus3 full particle in complex with 6D10 Fab
+Supramolecule #1: Echovirus3 full particle in complex with 6D10 Fab
+Supramolecule #3: 6D10 Fab
+Supramolecule #2: Echovirus E3
+Macromolecule #1: Heavy chain of 6D10
+Macromolecule #2: Light chain of 6D10
+Macromolecule #3: Genome polyprotein (Fragment)
+Macromolecule #4: Genome polyprotein
+Macromolecule #5: Genome polyprotein
+Macromolecule #6: Genome polyprotein (Fragment)
+Macromolecule #7: SPHINGOSINE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 65284 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X