[English] 日本語
 Yorodumi
Yorodumi- EMDB-33896: Structure of TRPA1 in Drosophila melanogaster in a state with 5 a... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of TRPA1 in Drosophila melanogaster in a state with 5 ankyrin repeats determined | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | ion channel / TRPA1 / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcellular response to pulsatile fluid shear stress / detection of hot stimulus involved in thermoception / temperature compensation of the circadian clock / negative phototaxis / TRP channels / neuronal signal transduction / thermosensory behavior / temperature-gated cation channel activity / mechanosensory behavior / detection of chemical stimulus involved in sensory perception of bitter taste ...cellular response to pulsatile fluid shear stress / detection of hot stimulus involved in thermoception / temperature compensation of the circadian clock / negative phototaxis / TRP channels / neuronal signal transduction / thermosensory behavior / temperature-gated cation channel activity / mechanosensory behavior / detection of chemical stimulus involved in sensory perception of bitter taste / cation channel complex / detection of chemical stimulus involved in sensory perception of pain / thermotaxis / detection of temperature stimulus involved in thermoception / monoatomic cation transmembrane transport / monoatomic cation transport / detection of temperature stimulus involved in sensory perception of pain / phototransduction / ligand-gated monoatomic ion channel activity / response to light stimulus / monoatomic cation channel activity / positive regulation of calcium-mediated signaling / calcium channel activity / calcium ion transport / cellular response to heat / response to heat / membrane / plasma membrane Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
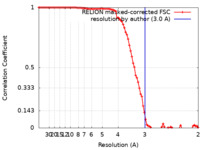
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||||||||
 Authors Authors | Sun L / Liu X / Yang Z / Wang X | |||||||||||||||
| Funding support |  China, 4 items China, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2023 Journal: Cell Discov / Year: 2023Title: Molecular architecture and gating mechanisms of the Drosophila TRPA1 channel. Authors: Xiaofei Wang / Yawen Li / Hong Wei / Zhisen Yang / Rui Luo / Yongxiang Gao / Wei Zhang / Xin Liu / Linfeng Sun /  Abstract: The transient receptor potential channel subfamily A member 1 (TRPA1) ion channel is an evolutionary conserved polymodal sensor responding to noxious temperature or chemical stimuli. Notably, the ...The transient receptor potential channel subfamily A member 1 (TRPA1) ion channel is an evolutionary conserved polymodal sensor responding to noxious temperature or chemical stimuli. Notably, the thermosensitivity of TRPA1 varies among different species or even different isoforms in the same species. However, the underlying molecular basis of its thermo-gating remains largely unknown. Here, we determine the structures of a heat-sensitive isoform of TRPA1 in Drosophila melanogaster in two distinct conformations with cryo-samples prepared at 8 °C. Large conformational changes are observed in the ankyrin repeat domain (ARD) and the coiled-coil domain between the two states. Remarkably, all 17 ankyrin repeats are mapped in the newly resolved conformation, forming a propeller-like architecture. Two intersubunit interfaces are identified in the amino (N)-terminal domain, and play vital roles during both heat and chemical activation as shown by electrophysiological analysis. With cryo-samples prepared at 35 °C, only one conformation is resolved, suggesting possible state transitions during heat responses. These findings provide a basis for further understanding how the ARD regulates channel functions, and insights into the gating mechanism of TRPA1. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33896.map.gz emd_33896.map.gz | 33.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33896-v30.xml emd-33896-v30.xml emd-33896.xml emd-33896.xml | 14.7 KB 14.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33896_fsc.xml emd_33896_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_33896.png emd_33896.png | 122.4 KB | ||
| Filedesc metadata |  emd-33896.cif.gz emd-33896.cif.gz | 6 KB | ||
| Others |  emd_33896_half_map_1.map.gz emd_33896_half_map_1.map.gz emd_33896_half_map_2.map.gz emd_33896_half_map_2.map.gz | 58.9 MB 58.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33896 http://ftp.pdbj.org/pub/emdb/structures/EMD-33896 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33896 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33896 | HTTPS FTP |
-Validation report
| Summary document |  emd_33896_validation.pdf.gz emd_33896_validation.pdf.gz | 845.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33896_full_validation.pdf.gz emd_33896_full_validation.pdf.gz | 844.6 KB | Display | |
| Data in XML |  emd_33896_validation.xml.gz emd_33896_validation.xml.gz | 16.5 KB | Display | |
| Data in CIF |  emd_33896_validation.cif.gz emd_33896_validation.cif.gz | 21.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33896 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33896 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33896 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33896 | HTTPS FTP |
-Related structure data
| Related structure data |  7yksMC  7ykrC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33896.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33896.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.01 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_33896_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_33896_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Homotetramer of dTRPA1
| Entire | Name: Homotetramer of dTRPA1 |
|---|---|
| Components |
|
-Supramolecule #1: Homotetramer of dTRPA1
| Supramolecule | Name: Homotetramer of dTRPA1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Transient receptor potential cation channel subfamily A member 1
| Macromolecule | Name: Transient receptor potential cation channel subfamily A member 1 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 135.324359 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MTSGDKETPK REDFASALRF LMGGCAREPE MTAMAPLNLP KKWARILRMS STPKIPIVDY LEAAESGNLD DFKRLFMADN SRIALKDAK GRTAAHQAAA RNRVNILRYI RDQNGDFNAK DNAGNTPLHI AVESDAYDAL DYLLSIPVDT GVLNEKKQAP V HLATELNK ...String: MTSGDKETPK REDFASALRF LMGGCAREPE MTAMAPLNLP KKWARILRMS STPKIPIVDY LEAAESGNLD DFKRLFMADN SRIALKDAK GRTAAHQAAA RNRVNILRYI RDQNGDFNAK DNAGNTPLHI AVESDAYDAL DYLLSIPVDT GVLNEKKQAP V HLATELNK VKSLRVMGQY RNVIDIQQGG EHGRTALHLA AIYDHEECAR ILITEFDACP RKPCNNGYYP IHEAAKNASS KT MEVFFQW GEQRGCTREE MISFYDSEGN VPLHSAVHGG DIKAVELCLK SGAKISTQQH DLSTPVHLAC AQGAIDIVKL MFE MQPMEK RLCLSCTDVQ KMTPLHCASM FDHPDIVSYL VAEGADINAL DKEHRSPLLL AASRSGWKTV HLLIRLGACI SVKD AAARN VLHFVIMNGG RLTDFAEQVA NCQTQAQLKL LLNEKDSMGC SPLHYASRDG HIRSLENLIR LGACINLKNN NNESP LHFA ARYGRYNTVR QLLDSEKGSF IINESDGAGM TPLHISSQQG HTRVVQLLLN RGALLHRDHT GRNPLQLAAM SGYTET IEL LHSVHSHLLD QVDKDGNTAL HLATMENKPH AISVLMSMGC KLVYNVLDMS AIDYAIYYKY PEAALAMVTH EERANEV MA LRSDKHPCVT LALIASMPKV FEAVQDKCIT KANCKKDSKS FYIKYSFAFL QCPFMFAKID EKTGESITTA SPIPLPAL N TMVTHGRVEL LAHPLSQKYL QMKWNSYGKY FHLANLLIYS IFLVFVTIYS SLMMNNIELK AGDNKTMSQY CNMGWEQLT MNLSQNPSVA SQIRLDSCEE RINRTTAILF CAVVIVVYIL LNSMRELIQI YQQKLHYILE TVNLISWVLY ISALVMVTPA FQPDGGINT IHYSAASIAV FLSWFRLLLF LQRFDQVGIY VVMFLEILQT LIKVLMVFSI LIIAFGLAFY ILLSKIIDPQ P NHLSFSNI PMSLLRTFSM MLGELDFVGT YVNTYYRDQL KVPMTSFLIL SVFMILMPIL LMNLLIGLAV GDIESVRRNA QL KRLAMQV VLHTELERKL PHVWLQRVDK MELIEYPNET KCKLGFCDFI LRKWFSNPFT EDSSMDVISF DNNDDYINAE LER QRRKLR DISRMLEQQH HLVRLIVQKM EIKTEADDVD EGISPNELRS VVGLRSAGGN RWNSPRVRNK LRAALSFNKS M UniProtKB: Transient receptor potential cation channel subfamily A member 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: DIFFRACTION / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)