+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of human PS2-containing gamma-secretase | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of calcium import into the mitochondrion / : / amyloid precursor protein biosynthetic process / gamma-secretase complex / aspartic endopeptidase activity, intramembrane cleaving / short-term synaptic potentiation / positive regulation of amyloid precursor protein biosynthetic process / positive regulation of endopeptidase activity / Noncanonical activation of NOTCH3 / Notch receptor processing ...regulation of calcium import into the mitochondrion / : / amyloid precursor protein biosynthetic process / gamma-secretase complex / aspartic endopeptidase activity, intramembrane cleaving / short-term synaptic potentiation / positive regulation of amyloid precursor protein biosynthetic process / positive regulation of endopeptidase activity / Noncanonical activation of NOTCH3 / Notch receptor processing / central nervous system myelination / membrane protein intracellular domain proteolysis / NOTCH4 Activation and Transmission of Signal to the Nucleus / growth factor receptor binding / Regulated proteolysis of p75NTR / metanephros development / glutamate receptor signaling pathway / amyloid precursor protein metabolic process / regulation of long-term synaptic potentiation / nuclear inner membrane / ciliary rootlet / myeloid cell homeostasis / azurophil granule membrane / G protein-coupled dopamine receptor signaling pathway / Hydrolases; Acting on peptide bonds (peptidases); Aspartic endopeptidases / Golgi cisterna membrane / adult behavior / mitochondrion-endoplasmic reticulum membrane tethering / positive regulation of catalytic activity / amyloid precursor protein catabolic process / plasma membrane => GO:0005886 / amyloid-beta formation / membrane protein ectodomain proteolysis / endopeptidase activator activity / EPH-ephrin mediated repulsion of cells / amyloid-beta metabolic process / T cell proliferation / Nuclear signaling by ERBB4 / Notch signaling pathway / NOTCH2 Activation and Transmission of Signal to the Nucleus / cellular response to calcium ion / Degradation of the extracellular matrix / NRIF signals cell death from the nucleus / Activated NOTCH1 Transmits Signal to the Nucleus / cerebellum development / dendritic shaft / epithelial cell proliferation / NOTCH3 Activation and Transmission of Signal to the Nucleus / neuromuscular junction / protein processing / sarcolemma / kinetochore / Constitutive Signaling by NOTCH1 PEST Domain Mutants / Constitutive Signaling by NOTCH1 HD+PEST Domain Mutants / Z disc / calcium ion transport / melanosome / synaptic vesicle / protein-macromolecule adaptor activity / presynaptic membrane / cell cortex / growth cone / ATPase binding / neuron apoptotic process / endopeptidase activity / mitochondrial inner membrane / learning or memory / early endosome / endosome membrane / response to hypoxia / intracellular signal transduction / membrane raft / apical plasma membrane / Amyloid fiber formation / lysosomal membrane / Golgi membrane / focal adhesion / centrosome / neuronal cell body / Neutrophil degranulation / endoplasmic reticulum membrane / negative regulation of apoptotic process / perinuclear region of cytoplasm / Golgi apparatus / enzyme binding / cell surface / endoplasmic reticulum / protein-containing complex / proteolysis / extracellular exosome / membrane / nucleus / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Guo X / Wang Y / Zhou J / Jin C / Wang J / Jia B / Jing D / Yan C / Lei J / Zhou R / Shi Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Molecular basis for isoform-selective inhibition of presenilin-1 by MRK-560. Authors: Xuefei Guo / Yumeng Wang / Jiayao Zhou / Chen Jin / Jiaoni Wang / Bojun Jia / Dan Jing / Chuangye Yan / Jianlin Lei / Rui Zhou / Yigong Shi /  Abstract: Inhibition of γ-secretase activity represents a potential therapeutic strategy for Alzheimer's disease (AD). MRK-560 is a selective inhibitor with higher potency for Presenilin 1 (PS1) than for PS2, ...Inhibition of γ-secretase activity represents a potential therapeutic strategy for Alzheimer's disease (AD). MRK-560 is a selective inhibitor with higher potency for Presenilin 1 (PS1) than for PS2, the two isoforms of the catalytic subunit of γ-secretase, although the underlying mechanism remains elusive. Here we report the cryo-electron microscopy (cryo-EM) structures of PS1 and PS2-containing γ-secretase complexes with and without MRK-560 at overall resolutions of 2.9-3.4 Å. MRK-560 occupies the substrate binding site of PS1, but is invisible in PS2. Structural comparison identifies Thr281 and Leu282 in PS1 to be the determinant for isoform-dependent sensitivity to MRK-560, which is confirmed by swapping experiment between PS1 and PS2. By revealing the mechanism for isoform-selective inhibition of presenilin, our work may facilitate future drug discovery targeting γ-secretase. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33629.map.gz emd_33629.map.gz | 28.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33629-v30.xml emd-33629-v30.xml emd-33629.xml emd-33629.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33629.png emd_33629.png | 30 KB | ||
| Others |  emd_33629_half_map_1.map.gz emd_33629_half_map_1.map.gz emd_33629_half_map_2.map.gz emd_33629_half_map_2.map.gz | 23.4 MB 23.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33629 http://ftp.pdbj.org/pub/emdb/structures/EMD-33629 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33629 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33629 | HTTPS FTP |
-Validation report
| Summary document |  emd_33629_validation.pdf.gz emd_33629_validation.pdf.gz | 626.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33629_full_validation.pdf.gz emd_33629_full_validation.pdf.gz | 625.9 KB | Display | |
| Data in XML |  emd_33629_validation.xml.gz emd_33629_validation.xml.gz | 10.6 KB | Display | |
| Data in CIF |  emd_33629_validation.cif.gz emd_33629_validation.cif.gz | 12.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33629 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33629 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33629 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33629 | HTTPS FTP |
-Related structure data
| Related structure data |  7y5zMC  7y5tC  7y5xC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33629.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33629.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.0825 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: CryoEM structure of human PS2-containing gamma-secretase
| File | emd_33629_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM structure of human PS2-containing gamma-secretase | ||||||||||||
| Projections & Slices |
| ||||||||||||
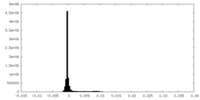
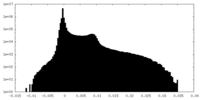
| Density Histograms |
-Half map: #1
| File | emd_33629_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
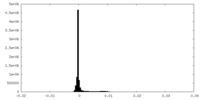
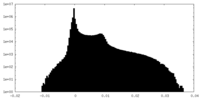
| Density Histograms |
- Sample components
Sample components
-Entire : Human PS2-containing gamma-secretase
| Entire | Name: Human PS2-containing gamma-secretase |
|---|---|
| Components |
|
-Supramolecule #1: Human PS2-containing gamma-secretase
| Supramolecule | Name: Human PS2-containing gamma-secretase / type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Nicastrin
| Macromolecule | Name: Nicastrin / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 78.48357 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MATAGGGSGA DPGSRGLLRL LSFCVLLAGL CRGNSVERKI YIPLNKTAPC VRLLNATHQI GCQSSISGDT GVIHVVEKEE DLQWVLTDG PNPPYMVLLE SKHFTRDLME KLKGRTSRIA GLAVSLTKPS PASGFSPSVQ CPNDGFGVYS NSYGPEFAHC R EIQWNSLG ...String: MATAGGGSGA DPGSRGLLRL LSFCVLLAGL CRGNSVERKI YIPLNKTAPC VRLLNATHQI GCQSSISGDT GVIHVVEKEE DLQWVLTDG PNPPYMVLLE SKHFTRDLME KLKGRTSRIA GLAVSLTKPS PASGFSPSVQ CPNDGFGVYS NSYGPEFAHC R EIQWNSLG NGLAYEDFSF PIFLLEDENE TKVIKQCYQD HNLSQNGSAP TFPLCAMQLF SHMHAVISTA TCMRRSSIQS TF SINPEIV CDPLSDYNVW SMLKPINTTG TLKPDDRVVV AATRLDSRSF FWNVAPGAES AVASFVTQLA AAEALQKAPD VTT LPRNVM FVFFQGETFD YIGSSRMVYD MEKGKFPVQL ENVDSFVELG QVALRTSLEL WMHTDPVSQK NESVRNQVED LLAT LEKSG AGVPAVILRR PNQSQPLPPS SLQRFLRARN ISGVVLADHS GAFHNKYYQS IYDTAENINV SYPEWLSPEE DLNFV TDTA KALADVATVL GRALYELAGG TNFSDTVQAD PQTVTRLLYG FLIKANNSWF QSILRQDLRS YLGDGPLQHY IAVSSP TNT TYVVQYALAN LTGTVVNLTR EQCQDPSKVP SENKDLYEYS WVQGPLHSNE TDRLPRCVRS TARLARALSP AFELSQW SS TEYSTWTESR WKDIRARIFL IASKELELIT LTVGFGILIF SLIVTYCINA KADVLFIAPR EPGAVSY |
-Macromolecule #2: Presenilin-2
| Macromolecule | Name: Presenilin-2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on peptide bonds (peptidases); Aspartic endopeptidases |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50.172035 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MLTFMASDSE EEVCDERTSL MSAESPTPRS CQEGRQGPED GENTAQWRSQ ENEEDGEEDP DRYVCSGVPG RPPGLEEELT LKYGAKHVI MLFVPVTLCM IVVVATIKSV RFYTEKNGQL IYTPFTEDTP SVGQRLLNSV LNTLIMISVI VVMTIFLVVL Y KYRCYKFI ...String: MLTFMASDSE EEVCDERTSL MSAESPTPRS CQEGRQGPED GENTAQWRSQ ENEEDGEEDP DRYVCSGVPG RPPGLEEELT LKYGAKHVI MLFVPVTLCM IVVVATIKSV RFYTEKNGQL IYTPFTEDTP SVGQRLLNSV LNTLIMISVI VVMTIFLVVL Y KYRCYKFI HGWLIMSSLM LLFLFTYIYL GEVLKTYNVA MDYPTLLLTV WNFGAVGMVC IHWKGPLVLQ QAYLIMISAL MA LVFIKYL PEWSAWVILG AISVYDLVAV LCPKGPLRML VETAQERNEP IFPALIYSSA MVWTVGMAKL DPSSQGALQL PYD PEMEED SYDSFGEPSY PEVFEPPLTG YPGEELEEEE ERGVKLGLGD FIFYSVLVGK AAATGSGDWN TTLACFVAIL IGLC LTLLL LAVFKKALPA LPISITFGLI FYFSTDNLVR PFMDTLASHQ LYI |
-Macromolecule #3: Gamma-secretase subunit APH-1A
| Macromolecule | Name: Gamma-secretase subunit APH-1A / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 29.017943 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGAAVFFGCT FVAFGPAFAL FLITVAGDPL RVIILVAGAF FWLVSLLLAS VVWFILVHVT DRSDARLQYG LLIFGAAVSV LLQEVFRFA YYKLLKKADE GLASLSEDGR SPISIRQMAY VSGLSFGIIS GVFSVINILA DALGPGVVGI HGDSPYYFLT S AFLTAAII ...String: MGAAVFFGCT FVAFGPAFAL FLITVAGDPL RVIILVAGAF FWLVSLLLAS VVWFILVHVT DRSDARLQYG LLIFGAAVSV LLQEVFRFA YYKLLKKADE GLASLSEDGR SPISIRQMAY VSGLSFGIIS GVFSVINILA DALGPGVVGI HGDSPYYFLT S AFLTAAII LLHTFWGVVF FDACERRRYW ALGLVVGSHL LTSGLTFLNP WYEASLLPIY AVTVSMGLWA FITAGGSLRS IQ RSLLCRR QEDSRVMVYS ALRIPPED |
-Macromolecule #4: Gamma-secretase subunit PEN-2
| Macromolecule | Name: Gamma-secretase subunit PEN-2 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.038029 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MNLERVSNEE KLNLCRKYYL GGFAFLPFLW LVNIFWFFRE AFLVPAYTEQ SQIKGYVWRS AVGFLFWVIV LTSWITIFQI YRPRWGALG DYLSFTIPLG TP |
-Macromolecule #7: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 7 / Number of copies: 6 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #8: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 8 / Number of copies: 1 / Formula: PC1 |
|---|---|
| Molecular weight | Theoretical: 790.145 Da |
| Chemical component information |  ChemComp-PC1: |
-Macromolecule #9: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 9 / Number of copies: 3 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.4 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 163958 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller

















 Z
Z Y
Y X
X