[English] 日本語
 Yorodumi
Yorodumi- EMDB-33219: Cryo-EM reconstruction of partial transmembrane channel E289A mut... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM reconstruction of partial transmembrane channel E289A mutant Vibrio cholerae Cytolysin | |||||||||
 Map data Map data | Cryo-EM reconstruction of partial transmembrane channel E289A mutant Vibrio cholerae Cytolysin | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
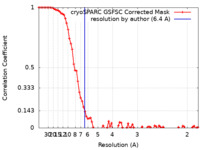
| Method | single particle reconstruction / cryo EM / Resolution: 6.4 Å | |||||||||
 Authors Authors | Mondal AK / Sengupta N / Singh M / Lata K / Lahiri I / Dutta S / Chattopadhyay K | |||||||||
| Funding support |  India, 2 items India, 2 items
| |||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2022 Journal: J Biol Chem / Year: 2022Title: Glu289 residue in the pore-forming motif of Vibrio cholerae cytolysin is important for efficient β-barrel pore formation. Authors: Anish Kumar Mondal / Nayanika Sengupta / Mahendra Singh / Rupam Biswas / Kusum Lata / Indrajit Lahiri / Somnath Dutta / Kausik Chattopadhyay /  Abstract: Vibrio cholerae cytolysin (VCC) is a potent membrane-damaging β-barrel pore-forming toxin. Upon binding to the target membranes, VCC monomers first assemble into oligomeric prepore intermediates and ...Vibrio cholerae cytolysin (VCC) is a potent membrane-damaging β-barrel pore-forming toxin. Upon binding to the target membranes, VCC monomers first assemble into oligomeric prepore intermediates and subsequently transform into transmembrane β-barrel pores. VCC harbors a designated pore-forming motif, which, during oligomeric pore formation, inserts into the membrane and generates a transmembrane β-barrel scaffold. It remains an enigma how the molecular architecture of the pore-forming motif regulates the VCC pore-formation mechanism. Here, we show that a specific pore-forming motif residue, E289, plays crucial regulatory roles in the pore-formation mechanism of VCC. We find that the mutation of E289A drastically compromises pore-forming activity, without affecting the structural integrity and membrane-binding potential of the toxin monomers. Although our single-particle cryo-EM analysis reveals WT-like oligomeric β-barrel pore formation by E289A-VCC in the membrane, we demonstrate that the mutant shows severely delayed kinetics in terms of pore-forming ability that can be rescued with elevated temperature conditions. We find that the pore-formation efficacy of E289A-VCC appears to be more profoundly dependent on temperature than that of the WT toxin. Our results suggest that the E289A mutation traps membrane-bound toxin molecules in the prepore-like intermediate state that is hindered from converting into the functional β-barrel pores by a large energy barrier, thus highlighting the importance of this residue for the pore-formation mechanism of VCC. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33219.map.gz emd_33219.map.gz | 44.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33219-v30.xml emd-33219-v30.xml emd-33219.xml emd-33219.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33219_fsc.xml emd_33219_fsc.xml | 8.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_33219.png emd_33219.png | 47.3 KB | ||
| Others |  emd_33219_half_map_1.map.gz emd_33219_half_map_1.map.gz emd_33219_half_map_2.map.gz emd_33219_half_map_2.map.gz | 44.2 MB 44.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33219 http://ftp.pdbj.org/pub/emdb/structures/EMD-33219 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33219 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33219 | HTTPS FTP |
-Validation report
| Summary document |  emd_33219_validation.pdf.gz emd_33219_validation.pdf.gz | 820.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33219_full_validation.pdf.gz emd_33219_full_validation.pdf.gz | 820.4 KB | Display | |
| Data in XML |  emd_33219_validation.xml.gz emd_33219_validation.xml.gz | 15.5 KB | Display | |
| Data in CIF |  emd_33219_validation.cif.gz emd_33219_validation.cif.gz | 19.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33219 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33219 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33219 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33219 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33219.map.gz / Format: CCP4 / Size: 47.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33219.map.gz / Format: CCP4 / Size: 47.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM reconstruction of partial transmembrane channel E289A mutant Vibrio cholerae Cytolysin | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.92 Å | ||||||||||||||||||||||||||||||||||||
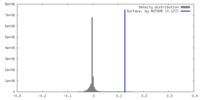
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half-map 1 of partial transmembrane channel E289A mutant...
| File | emd_33219_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 1 of partial transmembrane channel E289A mutant Vibrio cholerae Cytolysin | ||||||||||||
| Projections & Slices |
| ||||||||||||
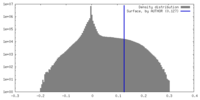
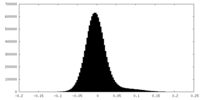
| Density Histograms |
-Half map: Half-map 2 of partial transmembrane channel E289A mutant...
| File | emd_33219_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 2 of partial transmembrane channel E289A mutant Vibrio cholerae Cytolysin | ||||||||||||
| Projections & Slices |
| ||||||||||||
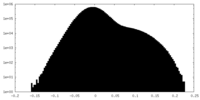
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM reconstruction of partial transmembrane channel E289A mut...
| Entire | Name: Cryo-EM reconstruction of partial transmembrane channel E289A mutant Vibrio cholerae Cytolysin |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM reconstruction of partial transmembrane channel E289A mut...
| Supramolecule | Name: Cryo-EM reconstruction of partial transmembrane channel E289A mutant Vibrio cholerae Cytolysin type: complex / Chimera: Yes / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.75 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)