+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
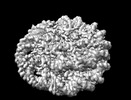
| タイトル | NuA4 bound to the nucleosome | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報PI5P Regulates TP53 Acetylation / RHOB GTPase cycle / NuA3b histone acetyltransferase complex / NuA3 histone acetyltransferase complex / NuA3a histone acetyltransferase complex / RHOA GTPase cycle / DNA Damage/Telomere Stress Induced Senescence / Sensing of DNA Double Strand Breaks / cellular bud neck contractile ring / piccolo histone acetyltransferase complex ...PI5P Regulates TP53 Acetylation / RHOB GTPase cycle / NuA3b histone acetyltransferase complex / NuA3 histone acetyltransferase complex / NuA3a histone acetyltransferase complex / RHOA GTPase cycle / DNA Damage/Telomere Stress Induced Senescence / Sensing of DNA Double Strand Breaks / cellular bud neck contractile ring / piccolo histone acetyltransferase complex / ascospore wall assembly / mitotic actomyosin contractile ring contraction / vacuole inheritance / histone H4K16 acetyltransferase activity / peptide 2-hydroxyisobutyryltransferase activity / histone crotonyltransferase activity / SUMOylation of transcription cofactors / DNA-templated transcription elongation / positive regulation of triglyceride biosynthetic process / actin cortical patch / SLIK (SAGA-like) complex / Swr1 complex / histone H4 acetyltransferase activity / rDNA heterochromatin formation / kinetochore assembly / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / peptide N-acetyltransferase activity / Ino80 complex / SWI/SNF complex / SAGA complex / peptide-lysine-N-acetyltransferase activity / establishment of cell polarity / NuA4 histone acetyltransferase complex / actin filament bundle / Estrogen-dependent gene expression / positive regulation of macroautophagy / protein secretion / chromosome organization / Ub-specific processing proteases / meiotic cell cycle / positive regulation of transcription elongation by RNA polymerase II / histone acetyltransferase activity / histone acetyltransferase / transcription coregulator activity / 転移酵素; アシル基を移すもの; アミノアシル基以外のアシル基を移すもの / methylated histone binding / actin filament / 加水分解酵素; 酸無水物に作用; 酸無水物に作用・細胞または細胞小器官の運動に関与 / structural constituent of cytoskeleton / structural constituent of chromatin / endocytosis / transcription corepressor activity / nucleosome / chromatin organization / histone binding / protein-containing complex assembly / chromatin remodeling / regulation of cell cycle / protein heterodimerization activity / DNA repair / chromatin binding / DNA-templated transcription / regulation of transcription by RNA polymerase II / chromatin / regulation of DNA-templated transcription / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / ATP hydrolysis activity / DNA binding / ATP binding / identical protein binding / nucleus / metal ion binding / cytosol 類似検索 - 分子機能 | |||||||||
| 生物種 |   | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 8.8 Å | |||||||||
 データ登録者 データ登録者 | Qu K / Chen Z | |||||||||
| 資金援助 | 1件
| |||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2022 ジャーナル: Nature / 年: 2022タイトル: Structure of the NuA4 acetyltransferase complex bound to the nucleosome. 著者: Keke Qu / Kangjing Chen / Hao Wang / Xueming Li / Zhucheng Chen /  要旨: Deoxyribonucleic acid in eukaryotes wraps around the histone octamer to form nucleosomes, the fundamental unit of chromatin. The N termini of histone H4 interact with nearby nucleosomes and play an ...Deoxyribonucleic acid in eukaryotes wraps around the histone octamer to form nucleosomes, the fundamental unit of chromatin. The N termini of histone H4 interact with nearby nucleosomes and play an important role in the formation of high-order chromatin structure and heterochromatin silencing. NuA4 in yeast and its homologue Tip60 complex in mammalian cells are the key enzymes that catalyse H4 acetylation, which in turn regulates chromatin packaging and function in transcription activation and DNA repair. Here we report the cryo-electron microscopy structure of NuA4 from Saccharomyces cerevisiae bound to the nucleosome. NuA4 comprises two major modules: the catalytic histone acetyltransferase (HAT) module and the transcription activator-binding (TRA) module. The nucleosome is mainly bound by the HAT module and is positioned close to a polybasic surface of the TRA module, which is important for the optimal activity of NuA4. The nucleosomal linker DNA carrying the upstream activation sequence is oriented towards the conserved, transcription activator-binding surface of the Tra1 subunit, which suggests a potential mechanism of NuA4 to act as a transcription co-activator. The HAT module recognizes the disk face of the nucleosome through the H2A-H2B acidic patch and nucleosomal DNA, projecting the catalytic pocket of Esa1 to the N-terminal tail of H4 and supporting its function in selective acetylation of H4. Together, our findings illustrate how NuA4 is assembled and provide mechanistic insights into nucleosome recognition and transcription co-activation by a HAT. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_32150.map.gz emd_32150.map.gz | 2.2 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-32150-v30.xml emd-32150-v30.xml emd-32150.xml emd-32150.xml | 33.7 KB 33.7 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_32150.png emd_32150.png | 28.7 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32150 http://ftp.pdbj.org/pub/emdb/structures/EMD-32150 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32150 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32150 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7vvzMC  7vvuC  7vvyC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_32150.map.gz / 形式: CCP4 / 大きさ: 3.2 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_32150.map.gz / 形式: CCP4 / 大きさ: 3.2 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ボクセルのサイズ | X=Y=Z: 4.33 Å | ||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : NuA4 bound to the nucleosome
+超分子 #1: NuA4 bound to the nucleosome
+分子 #1: Chromatin modification-related protein EAF6
+分子 #2: Chromatin modification-related protein YNG2
+分子 #3: Enhancer of polycomb-like protein 1
+分子 #4: Histone H3
+分子 #5: Histone H4
+分子 #6: Histone H2A
+分子 #7: Histone H2B 1.1
+分子 #8: Histone acetyltransferase ESA1
+分子 #11: Epl1 arginine anchor
+分子 #12: Chromatin modification-related protein EAF1
+分子 #13: Actin-related protein 4
+分子 #14: Actin
+分子 #15: SWR1-complex protein 4
+分子 #16: Transcription-associated protein 1
+分子 #9: DNA (207-mer)
+分子 #10: DNA (207-mer)
+分子 #17: CARBOXYMETHYL COENZYME *A
+分子 #18: MAGNESIUM ION
+分子 #19: ADENOSINE-5'-TRIPHOSPHATE
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.6 |
|---|---|
| 凍結 | 凍結剤: NITROGEN |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 50.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 1.8 µm / 最小 デフォーカス(公称値): 1.3 µm |
- 画像解析
画像解析
| 最終 再構成 | 解像度のタイプ: BY AUTHOR / 解像度: 8.8 Å / 解像度の算出法: FSC 0.143 CUT-OFF / 使用した粒子像数: 474949 |
|---|---|
| 初期 角度割当 | タイプ: ANGULAR RECONSTITUTION |
| 最終 角度割当 | タイプ: ANGULAR RECONSTITUTION |
 ムービー
ムービー コントローラー
コントローラー