[English] 日本語
 Yorodumi
Yorodumi- EMDB-32049: Streptomyces coelicolor zinc uptake regulator complexed with zinc... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Streptomyces coelicolor zinc uptake regulator complexed with zinc and DNA (dimer of dimers) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of secondary metabolite biosynthetic process / DNA-binding transcription repressor activity / protein-DNA complex / transcription cis-regulatory region binding / DNA-binding transcription factor activity / negative regulation of DNA-templated transcription / zinc ion binding / cytosol Similarity search - Function | |||||||||
| Biological species |  Streptomyces coelicolor A3(2) (bacteria) / Streptomyces coelicolor A3(2) (bacteria) /  Streptomyces coelicolor (strain ATCC BAA-471 / A3(2) / M145) (bacteria) Streptomyces coelicolor (strain ATCC BAA-471 / A3(2) / M145) (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Yang X / Zheng J | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2022 Journal: Nucleic Acids Res / Year: 2022Title: Structural basis of Streptomyces transcription activation by zinc uptake regulator. Authors: Xu Yang / Yiqun Wang / Guiyang Liu / Zixin Deng / Shuangjun Lin / Jianting Zheng /  Abstract: Streptomyces coelicolor (Sc) is a model organism of actinobacteria to study morphological differentiation and production of bioactive metabolites. Sc zinc uptake regulator (Zur) affects both ...Streptomyces coelicolor (Sc) is a model organism of actinobacteria to study morphological differentiation and production of bioactive metabolites. Sc zinc uptake regulator (Zur) affects both processes by controlling zinc homeostasis. It activates transcription by binding to palindromic Zur-box sequences upstream of -35 elements. Here we deciphered the molecular mechanism by which ScZur interacts with promoter DNA and Sc RNA polymerase (RNAP) by cryo-EM structures and biochemical assays. The ScZur-DNA structures reveal a sequential and cooperative binding of three ScZur dimers surrounding a Zur-box spaced 8 nt upstream from a -35 element. The ScRNAPσHrdB-Zur-DNA structures define protein-protein and protein-DNA interactions involved in the principal housekeeping σHrdB-dependent transcription initiation from a noncanonical promoter with a -10 element lacking the critical adenine residue at position -11 and a TTGCCC -35 element deviating from the canonical TTGACA motif. ScZur interacts with the C-terminal domain of ScRNAP α subunit (αCTD) in a complex structure trapped in an active conformation. Key ScZur-αCTD interfacial residues accounting for ScZur-dependent transcription activation were confirmed by mutational studies. Together, our structural and biochemical results provide a comprehensive model for transcription activation of Zur family regulators. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32049.map.gz emd_32049.map.gz | 1.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32049-v30.xml emd-32049-v30.xml emd-32049.xml emd-32049.xml | 17.8 KB 17.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_32049.png emd_32049.png | 62.2 KB | ||
| Masks |  emd_32049_msk_1.map emd_32049_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-32049.cif.gz emd-32049.cif.gz | 5.7 KB | ||
| Others |  emd_32049_half_map_1.map.gz emd_32049_half_map_1.map.gz emd_32049_half_map_2.map.gz emd_32049_half_map_2.map.gz | 95.7 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32049 http://ftp.pdbj.org/pub/emdb/structures/EMD-32049 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32049 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32049 | HTTPS FTP |
-Validation report
| Summary document |  emd_32049_validation.pdf.gz emd_32049_validation.pdf.gz | 615.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_32049_full_validation.pdf.gz emd_32049_full_validation.pdf.gz | 614.9 KB | Display | |
| Data in XML |  emd_32049_validation.xml.gz emd_32049_validation.xml.gz | 12.8 KB | Display | |
| Data in CIF |  emd_32049_validation.cif.gz emd_32049_validation.cif.gz | 15 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32049 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32049 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32049 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32049 | HTTPS FTP |
-Related structure data
| Related structure data |  7vo9MC  7vo0C  7vpdC  7vpzC  7x74C  7x75C  7x76C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_32049.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32049.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
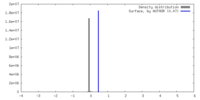
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_32049_msk_1.map emd_32049_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
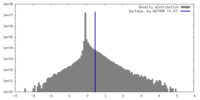
| Density Histograms |
-Half map: #2
| File | emd_32049_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
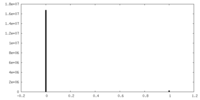
| Density Histograms |
-Half map: #1
| File | emd_32049_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
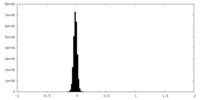
| Density Histograms |
- Sample components
Sample components
-Entire : Zur-DNA
| Entire | Name: Zur-DNA |
|---|---|
| Components |
|
-Supramolecule #1: Zur-DNA
| Supramolecule | Name: Zur-DNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Streptomyces coelicolor A3(2) (bacteria) Streptomyces coelicolor A3(2) (bacteria) |
| Molecular weight | Theoretical: 250 kDa/nm |
-Macromolecule #1: DNA (84-MER)
| Macromolecule | Name: DNA (84-MER) / type: dna / ID: 1 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Streptomyces coelicolor A3(2) (bacteria) Streptomyces coelicolor A3(2) (bacteria) |
| Molecular weight | Theoretical: 25.808424 KDa |
| Sequence | String: (DC)(DA)(DA)(DG)(DG)(DC)(DA)(DC)(DA)(DT) (DG)(DA)(DC)(DA)(DA)(DC)(DG)(DG)(DT)(DG) (DT)(DT)(DC)(DA)(DG)(DT)(DG)(DC)(DC) (DG)(DC)(DG)(DT)(DT)(DG)(DC)(DC)(DC)(DG) (DA) (DT)(DA)(DC)(DC)(DC)(DC) ...String: (DC)(DA)(DA)(DG)(DG)(DC)(DA)(DC)(DA)(DT) (DG)(DA)(DC)(DA)(DA)(DC)(DG)(DG)(DT)(DG) (DT)(DT)(DC)(DA)(DG)(DT)(DG)(DC)(DC) (DG)(DC)(DG)(DT)(DT)(DG)(DC)(DC)(DC)(DG) (DA) (DT)(DA)(DC)(DC)(DC)(DC)(DC)(DT) (DA)(DC)(DC)(DC)(DG)(DT)(DA)(DG)(DT)(DT) (DG)(DA) (DC)(DT)(DG)(DG)(DC)(DA)(DT) (DC)(DC)(DG)(DG)(DG)(DC)(DG)(DC)(DC)(DG) (DG)(DG)(DT) (DC)(DG)(DC)(DC) GENBANK: GENBANK: AL939129.1 |
-Macromolecule #2: DNA (84-MER)
| Macromolecule | Name: DNA (84-MER) / type: dna / ID: 2 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Streptomyces coelicolor A3(2) (bacteria) Streptomyces coelicolor A3(2) (bacteria) |
| Molecular weight | Theoretical: 26.008541 KDa |
| Sequence | String: (DG)(DG)(DC)(DG)(DA)(DC)(DC)(DC)(DG)(DG) (DC)(DG)(DC)(DC)(DG)(DC)(DC)(DT)(DA)(DC) (DG)(DG)(DT)(DC)(DA)(DG)(DT)(DA)(DC) (DT)(DA)(DC)(DG)(DG)(DG)(DT)(DA)(DG)(DG) (DG) (DG)(DG)(DT)(DA)(DT)(DC) ...String: (DG)(DG)(DC)(DG)(DA)(DC)(DC)(DC)(DG)(DG) (DC)(DG)(DC)(DC)(DG)(DC)(DC)(DT)(DA)(DC) (DG)(DG)(DT)(DC)(DA)(DG)(DT)(DA)(DC) (DT)(DA)(DC)(DG)(DG)(DG)(DT)(DA)(DG)(DG) (DG) (DG)(DG)(DT)(DA)(DT)(DC)(DG)(DG) (DG)(DC)(DA)(DA)(DC)(DG)(DC)(DG)(DG)(DC) (DA)(DC) (DT)(DG)(DA)(DA)(DC)(DA)(DC) (DC)(DG)(DT)(DT)(DG)(DT)(DC)(DA)(DT)(DG) (DT)(DG)(DC) (DC)(DT)(DT)(DG) |
-Macromolecule #3: Putative metal uptake regulation protein
| Macromolecule | Name: Putative metal uptake regulation protein / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Streptomyces coelicolor (strain ATCC BAA-471 / A3(2) / M145) (bacteria) Streptomyces coelicolor (strain ATCC BAA-471 / A3(2) / M145) (bacteria)Strain: ATCC BAA-471 / A3(2) / M145 |
| Molecular weight | Theoretical: 16.90484 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SSGLVPRGSH MTTAGPPVKG RATRQRAAVS AALQEVEEFR SAQELHDMLK HKGDAVGLTT VYRTLQSLAD AGEVDVLRT AEGESVYRRC STGDHHHHLV CRACGKAVEV EGPAVEKWAE AIAAEHGYVN VAHTVEIFGT CADCAGASGG UniProtKB: Metal uptake regulation protein |
-Macromolecule #4: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 4 / Number of copies: 12 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 289 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy #1
Electron microscopy #1
| Microscopy ID | 1 |
|---|---|
| Microscope | FEI TITAN KRIOS |
| Image recording | Image recording ID: 1 / Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 1.25 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Electron microscopy #1~
Electron microscopy #1~
| Microscopy ID | 1 |
|---|---|
| Microscope | FEI TITAN KRIOS |
| Image recording | Image recording ID: 2 / Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 1.25 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)