[English] 日本語
 Yorodumi
Yorodumi- EMDB-31077: Structure of Rift Valley fever virus RNA-dependent RNA polymerase -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31077 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Rift Valley fever virus RNA-dependent RNA polymerase | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationnucleoside binding / RNA-directed RNA polymerase / viral RNA genome replication / RNA-dependent RNA polymerase activity / DNA-templated transcription Similarity search - Function | |||||||||
| Biological species |   Rift Valley fever virus / Rift Valley fever virus /   Rift valley fever virus Rift valley fever virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Wang X / Hu CX | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: J Virol / Year: 2022 Journal: J Virol / Year: 2022Title: Structure of Rift Valley Fever Virus RNA-Dependent RNA Polymerase. Authors: Xue Wang / Cuixia Hu / Wei Ye / Jia Wang / Xiaofei Dong / Jie Xu / Xiaorong Li / Manfeng Zhang / Hongyun Lu / Fanglin Zhang / Wei Wu / Shaodong Dai / Hong-Wei Wang / Zhongzhou Chen /   Abstract: Rift Valley fever virus (RVFV) belongs to the order and is the type species of genus , which accounts for over 50% of family species. RVFV is mosquito-borne and causes severe diseases in both ...Rift Valley fever virus (RVFV) belongs to the order and is the type species of genus , which accounts for over 50% of family species. RVFV is mosquito-borne and causes severe diseases in both humans and livestock, and consists of three segments (S, M, L) in the genome. The L segment encodes an RNA-dependent RNA polymerase (RdRp, L protein) that is responsible for facilitating the replication and transcription of the virus. It is essential for the virus and has multiple drug targets. Here, we established an expression system and purification procedures for full-length L protein, which is composed of an endonuclease domain, RdRp domain, and cap-binding domain. A cryo-EM L protein structure was reported at 3.6 Å resolution. In this first L protein structure of genus , the priming loop of RVFV L protein is distinctly different from those of other L proteins and undergoes large movements related to its replication role. Structural and biochemical analyses indicate that a single template can induce initiation of RNA synthesis, which is notably enhanced by 5' viral RNA. These findings help advance our understanding of the mechanism of RNA synthesis and provide an important basis for developing antiviral inhibitors. The zoonosis RVF virus (RVFV) is one of the most serious arbovirus threats to both human and animal health. RNA-dependent RNA polymerase (RdRp) is a multifunctional enzyme catalyzing genome replication as well as viral transcription, so the RdRp is essential for studying the virus and has multiple drug targets. In our study, we report the structure of RVFV L protein at 3.6 Å resolution by cryo-EM. This is the first L protein structure of genus . Strikingly, a single template can initiate RNA replication. The structure and assays provide a comprehensive and in-depth understanding of the catalytic and substrate recognition mechanism of RdRp. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31077.map.gz emd_31077.map.gz | 2.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31077-v30.xml emd-31077-v30.xml emd-31077.xml emd-31077.xml | 11.4 KB 11.4 KB | Display Display |  EMDB header EMDB header |
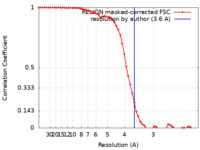
| FSC (resolution estimation) |  emd_31077_fsc.xml emd_31077_fsc.xml | 5.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_31077.png emd_31077.png | 19.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31077 http://ftp.pdbj.org/pub/emdb/structures/EMD-31077 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31077 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31077 | HTTPS FTP |
-Related structure data
| Related structure data |  7eeiMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_31077.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31077.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Structural insights into Rift Valley fever virus replication machinery
| Entire | Name: Structural insights into Rift Valley fever virus replication machinery |
|---|---|
| Components |
|
-Supramolecule #1: Structural insights into Rift Valley fever virus replication machinery
| Supramolecule | Name: Structural insights into Rift Valley fever virus replication machinery type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Rift Valley fever virus Rift Valley fever virus |
| Recombinant expression | Organism:  Pichia aff. alni PL5W1 (fungus) Pichia aff. alni PL5W1 (fungus) |
| Molecular weight | Experimental: 238 KDa |
-Macromolecule #1: Replicase
| Macromolecule | Name: Replicase / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: RNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:   Rift valley fever virus Rift valley fever virus |
| Molecular weight | Theoretical: 202.260734 KDa |
| Recombinant expression | Organism:  Pichia aff. alni PL5W1 (fungus) Pichia aff. alni PL5W1 (fungus) |
| Sequence | String: GKTERELLAM VSSIQINWSV TESVFPPFSR EMFDRFRSSP PDSEYITRIV SRCLINSQEK LINSSFFAEG NDKALRFSKN AEECSLAVE RALNQYRAED NLRDLNDHSS TIQLPPWLSY HDVDGKDLCP LQGLDVRGDH PMCNLWREVV TSANLEEIER M HDDAAAEL ...String: GKTERELLAM VSSIQINWSV TESVFPPFSR EMFDRFRSSP PDSEYITRIV SRCLINSQEK LINSSFFAEG NDKALRFSKN AEECSLAVE RALNQYRAED NLRDLNDHSS TIQLPPWLSY HDVDGKDLCP LQGLDVRGDH PMCNLWREVV TSANLEEIER M HDDAAAEL EFALSGVKDR PDERNRYHRV HLNMGSDDSV YIAALGVNGK KHKADTLVQQ MRDRSKQPFS PDHDVDHISE FL SACSSDL WATDEDLYNP LSCDKELRLA AQRIHQPSLS ERGFNEIITE HYKFMGSRIG SWCQMVSLIG AELSASVKQH VKP NYFVIK RLLGSGIFLL IKPTSSKSHI FVSFAIKRSC WAFDLSTSRV FKPYIDAGDL LVTDFVSYKL SKLTNLCKCV SLME SSFSF WAEAFGIPSW NFVGDLFRSS DSAAMDASYM GKLSLLTLLE DKAATEELQT IARYIIMEGF VSPPEIPKPH KMTSK FPKV LRSELQVYLL NCLCRTIQRI AGEPFILKKK DGSISWGGMF NPFSGRPLLD MQPLISCCYN GYFKNKEEET EPSSLS GMY KKIIELEHLR PQSDAFLGYK DPELPRMHEF SVSYLKEACN HAKLVLRSLY GQNFMEQIDN QIIRELSGLT LERLATL KA TSNFNENWYV YKDVADKNYT RDKLLVKMSK YASEGKSLAI QKFEDCMRQI ESQGCMHICL FKKQQHGGLR EIYVMGAE E RIVQSVVETI ARSIGKFFAS DTLCNPPNKV KIPETHGIRA RKQCKGPVWT CATSDDARKW NQGHFVTKFA LMLCEFTSP KWWPLIIRGC SMFTRKRMMM NLNYLKILDG HRELDIRDDF VMDLFKAYHG EAEVPWAFKG KTYLETTTGM MQGILHYTSS LLHTIHQEY IRSLSFKIFN LKVAPEMSKG LVCDMMQGSD DSSMLISFPA DDEKVLTRCK VAAAICFRMK KELGVYLAIY P SEKSTANT DFVMEYNSEF YFHTQHVRPT IRWIAACCSL PEVETLVARQ EEASNLMTSV TEGGGSFSLA AMIQQAQCTL HY MLMGMGV SELFLEYKKA VLKWNDPGLG FFLLDNPYAC GLGGFRFNLF KAITRTDLQK LYAFFMKKVK GSAARDWADE DVT IPETCS VSPGGALILS SSLKWGSRKK FQKLRDRLNI PENWIELINE NPEVLYRAPR TGPEILLRIA EKVHSPGVVS SLSS GNAVC KVMASAVYFL SATIFEDTGR PEFNFLEDSK YSLLQKMAAY SGFHGFNDME PEDILFLFPN IEELESLDSI VYNKG EIDI IPRVNIRDAT QTRVTIFNEQ KTLRTSPEKL VSDKWFGTQK SRIGKTTFLA EWEKLKKIVK WLEDTPEATL AHTPLN NHI QVRNFFARME SKPRTVRITG APVKKRSGVS KIAMVIRDNF SRMGHLRGVE DLAGFTRSVS AEILKHFLFC ILQGPYS ES YKLQLIYRVL SSVSNVEIKE SDGKTKTNLI GILQRFLDGD HVVPIIEEMG AGTVGGFIKR QQSKVVQNKV VYYGVGIW R GFMDGYQVHL EIENDIGQPP RLRNVTTNCQ SSPWDLSIPI RQWAEDMGVT NNQDYSSKSS RGARYWMHSF RMQGPSKPF GCPVYIIKGD MSDVIRLRKE EVEMKVRGST LNLYTKHHSH QDLHILSYTA SDNDLSPGIF KSISDEGVAQ ALQLFEREPS NCWVRCESV APKFISAILE ICEGKRQIRG INRTRLSEIV RICSESSLRS KVGSMFSFVA NVEEAHDVDY DALMDLMIED A KNNAFSHV VDCIELDV |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller