+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of WT E.coli 70S ribosome complexed with mRNA, P-site fMet-NH-tRNAfMet and A-site ortho-aminobenzoic acid charged NH-tRNAPhe | |||||||||
 マップデータ マップデータ | 70S global refined map | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | ribosome / non-natural monomers / aminobenzoic acids / unnatural monomers | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報negative regulation of cytoplasmic translational initiation / positive regulation of ribosome biogenesis / DnaA-L2 complex / negative regulation of translational initiation / negative regulation of DNA-templated DNA replication initiation / translational initiation / ribosome assembly / mRNA regulatory element binding translation repressor activity / assembly of large subunit precursor of preribosome / cytosolic ribosome assembly ...negative regulation of cytoplasmic translational initiation / positive regulation of ribosome biogenesis / DnaA-L2 complex / negative regulation of translational initiation / negative regulation of DNA-templated DNA replication initiation / translational initiation / ribosome assembly / mRNA regulatory element binding translation repressor activity / assembly of large subunit precursor of preribosome / cytosolic ribosome assembly / transcription antitermination / regulation of cell growth / DNA-templated transcription termination / maintenance of translational fidelity / ribosomal large subunit assembly / mRNA 5'-UTR binding / large ribosomal subunit / ribosome binding / ribosomal small subunit biogenesis / ribosomal small subunit assembly / small ribosomal subunit / small ribosomal subunit rRNA binding / transferase activity / 5S rRNA binding / large ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / cytosolic large ribosomal subunit / tRNA binding / cytoplasmic translation / rRNA binding / negative regulation of translation / ribosome / structural constituent of ribosome / ribonucleoprotein complex / translation / response to antibiotic / mRNA binding / RNA binding / zinc ion binding / membrane / cytoplasm / cytosol 類似検索 - 分子機能 | |||||||||
| 生物種 |  | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.02 Å | |||||||||
 データ登録者 データ登録者 | Majumdar C / Cate JHD | |||||||||
| 資金援助 |  米国, 1件 米国, 1件
| |||||||||
 引用 引用 |  ジャーナル: ACS Cent Sci / 年: 2023 ジャーナル: ACS Cent Sci / 年: 2023タイトル: Aminobenzoic Acid Derivatives Obstruct Induced Fit in the Catalytic Center of the Ribosome. 著者: Chandrima Majumdar / Joshua A Walker / Matthew B Francis / Alanna Schepartz / Jamie H D Cate /  要旨: The () ribosome can incorporate a variety of non-l-α-amino acid monomers into polypeptide chains but with poor efficiency. Although these monomers span a diverse set of compounds, there exists no ...The () ribosome can incorporate a variety of non-l-α-amino acid monomers into polypeptide chains but with poor efficiency. Although these monomers span a diverse set of compounds, there exists no high-resolution structural information regarding their positioning within the catalytic center of the ribosome, the peptidyl transferase center (PTC). Thus, details regarding the mechanism of amide bond formation and the structural basis for differences and defects in incorporation efficiency remain unknown. Within a set of three aminobenzoic acid derivatives-3-aminopyridine-4-carboxylic acid (Apy), aminobenzoic acid (ABZ), and aminobenzoic acid (ABZ)-the ribosome incorporates Apy into polypeptide chains with the highest efficiency, followed by ABZ and then ABZ, a trend that does not track with the nucleophilicity of the reactive amines. Here, we report high-resolution cryo-EM structures of the ribosome with each of these three aminobenzoic acid derivatives charged on tRNA bound in the aminoacyl-tRNA site (A-site). The structures reveal how the aromatic ring of each monomer sterically blocks the positioning of nucleotide U2506, thereby preventing rearrangement of nucleotide U2585 and the resulting induced fit in the PTC required for efficient amide bond formation. They also reveal disruptions to the bound water network that is believed to facilitate formation and breakdown of the tetrahedral intermediate. Together, the cryo-EM structures reported here provide a mechanistic rationale for differences in reactivity of aminobenzoic acid derivatives relative to l-α-amino acids and each other and identify stereochemical constraints on the size and geometry of non-monomers that can be accepted efficiently by wild-type ribosomes. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_29786.map.gz emd_29786.map.gz | 46 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-29786-v30.xml emd-29786-v30.xml emd-29786.xml emd-29786.xml | 74.2 KB 74.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_29786_fsc.xml emd_29786_fsc.xml | 16.3 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_29786.png emd_29786.png | 89.1 KB | ||
| Filedesc metadata |  emd-29786.cif.gz emd-29786.cif.gz | 14.8 KB | ||
| その他 |  emd_29786_half_map_1.map.gz emd_29786_half_map_1.map.gz emd_29786_half_map_2.map.gz emd_29786_half_map_2.map.gz | 305.1 MB 81.9 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29786 http://ftp.pdbj.org/pub/emdb/structures/EMD-29786 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29786 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29786 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_29786_validation.pdf.gz emd_29786_validation.pdf.gz | 866.7 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_29786_full_validation.pdf.gz emd_29786_full_validation.pdf.gz | 866.2 KB | 表示 | |
| XML形式データ |  emd_29786_validation.xml.gz emd_29786_validation.xml.gz | 24.2 KB | 表示 | |
| CIF形式データ |  emd_29786_validation.cif.gz emd_29786_validation.cif.gz | 32.1 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29786 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29786 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29786 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29786 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8g6wMC  8g6xC  8g6yC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_29786.map.gz / 形式: CCP4 / 大きさ: 381.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_29786.map.gz / 形式: CCP4 / 大きさ: 381.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | 70S global refined map | ||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.8232 Å | ||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: 70S global refinement half map
| ファイル | emd_29786_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | 70S global refinement half map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
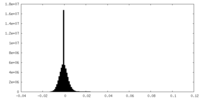
| 密度ヒストグラム |
-ハーフマップ: 70S global refinement half map
| ファイル | emd_29786_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | 70S global refinement half map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : 70S E.coli ribosome
+超分子 #1: 70S E.coli ribosome
+分子 #1: 50S ribosomal protein L33
+分子 #2: 50S ribosomal protein L34
+分子 #3: 50S ribosomal protein L35
+分子 #4: 50S ribosomal protein L36
+分子 #5: 50S ribosomal protein L31
+分子 #7: 30S ribosomal protein S2
+分子 #8: 30S ribosomal protein S3
+分子 #9: 30S ribosomal protein S4
+分子 #10: 30S ribosomal protein S5
+分子 #11: 30S ribosomal protein S6
+分子 #12: 30S ribosomal protein S7
+分子 #13: 30S ribosomal protein S8
+分子 #14: 30S ribosomal protein S9
+分子 #15: 30S ribosomal protein S10
+分子 #16: 30S ribosomal protein S11
+分子 #17: 30S ribosomal protein S12
+分子 #18: 30S ribosomal protein S13
+分子 #19: 30S ribosomal protein S14
+分子 #20: 30S ribosomal protein S15
+分子 #21: 30S ribosomal protein S16
+分子 #22: 30S ribosomal protein S17
+分子 #23: 30S ribosomal protein S18
+分子 #24: 30S ribosomal protein S19
+分子 #25: 30S ribosomal protein S20
+分子 #26: 30S ribosomal protein S21
+分子 #32: 50S ribosomal protein L2
+分子 #33: 50S ribosomal protein L3
+分子 #34: 50S ribosomal protein L4
+分子 #35: 50S ribosomal protein L5
+分子 #36: 50S ribosomal protein L6
+分子 #37: 50S ribosomal protein L9
+分子 #38: 50S ribosomal protein L13
+分子 #39: 50S ribosomal protein L14
+分子 #40: 50S ribosomal protein L15
+分子 #41: 50S ribosomal protein L16
+分子 #42: 50S ribosomal protein L17
+分子 #43: 50S ribosomal protein L18
+分子 #44: 50S ribosomal protein L19
+分子 #45: 50S ribosomal protein L20
+分子 #46: Ribosomal protein L21
+分子 #47: 50S ribosomal protein L22
+分子 #48: 50S ribosomal protein L23
+分子 #49: 50S ribosomal protein L24
+分子 #50: 50S ribosomal protein L25
+分子 #51: 50S ribosomal protein L27
+分子 #52: 50S ribosomal protein L28
+分子 #53: 50S ribosomal protein L29
+分子 #54: 50S ribosomal protein L30
+分子 #55: 50S ribosomal protein L32
+分子 #6: 16S rRNA
+分子 #27: mRNA
+分子 #28: A-site oABZ-tRNAPhe
+分子 #29: P-site fMet-tRNAfMet
+分子 #30: 23S rRNA
+分子 #31: 5S rRNA
+分子 #56: ZINC ION
+分子 #57: PAROMOMYCIN
+分子 #58: MAGNESIUM ION
+分子 #59: 2-AMINO-BENZAMIDE
+分子 #60: 3'-amino-3'-deoxyadenosine 5'-(dihydrogen phosphate)
+分子 #61: N-FORMYLMETHIONINE
+分子 #62: POTASSIUM ION
+分子 #63: SPERMIDINE
+分子 #64: water
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| グリッド | モデル: Quantifoil R1.2/1.3 / 材質: GOLD / メッシュ: 300 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: CONTINUOUS / 支持フィルム - Film thickness: 0.2 / 前処理 - タイプ: GLOW DISCHARGE |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277 K / 装置: FEI VITROBOT MARK IV |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 BIOQUANTUM (6k x 4k) 平均電子線量: 40.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: OTHER / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 1.5 µm / 最小 デフォーカス(公称値): 0.5 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー











 Z
Z Y
Y X
X