[English] 日本語
 Yorodumi
Yorodumi- EMDB-29728: Native GABA-A receptor from the mouse brain, alpha1-beta2-gamma2 ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Native GABA-A receptor from the mouse brain, alpha1-beta2-gamma2 subtype, in complex with didesethylflurazepam and endogenous GABA | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ligand-gated ion channel / cys-loop receptor / neurotransmitter receptor / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationGABA receptor activation / inhibitory extracellular ligand-gated monoatomic ion channel activity / diazepam binding / GABA receptor complex / organic cyclic compound binding / inner ear receptor cell development / cellular response to histamine / GABA-A receptor activity / GABA-gated chloride ion channel activity / GABA-A receptor complex ...GABA receptor activation / inhibitory extracellular ligand-gated monoatomic ion channel activity / diazepam binding / GABA receptor complex / organic cyclic compound binding / inner ear receptor cell development / cellular response to histamine / GABA-A receptor activity / GABA-gated chloride ion channel activity / GABA-A receptor complex / inhibitory synapse assembly / innervation / synaptic transmission, GABAergic / gamma-aminobutyric acid signaling pathway / postsynaptic specialization membrane / neurotransmitter receptor activity / chloride channel activity / cochlea development / chloride channel complex / regulation of postsynaptic membrane potential / transmembrane transporter complex / neuron development / GABA-ergic synapse / presynaptic active zone membrane / chloride transmembrane transport / dendrite membrane / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / cytoplasmic vesicle membrane / cytoplasmic vesicle / postsynapse / neuron projection / glutamatergic synapse / synapse / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
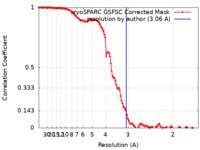
| Method | single particle reconstruction / cryo EM / Resolution: 3.06 Å | |||||||||
 Authors Authors | Sun C / Gouaux E | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation | Journal: Acta Crystallogr D Struct Biol / Year: 2018 Title: Real-space refinement in PHENIX for cryo-EM and crystallography. Authors: Pavel V Afonine / Billy K Poon / Randy J Read / Oleg V Sobolev / Thomas C Terwilliger / Alexandre Urzhumtsev / Paul D Adams /    Abstract: This article describes the implementation of real-space refinement in the phenix.real_space_refine program from the PHENIX suite. The use of a simplified refinement target function enables very fast ...This article describes the implementation of real-space refinement in the phenix.real_space_refine program from the PHENIX suite. The use of a simplified refinement target function enables very fast calculation, which in turn makes it possible to identify optimal data-restraint weights as part of routine refinements with little runtime cost. Refinement of atomic models against low-resolution data benefits from the inclusion of as much additional information as is available. In addition to standard restraints on covalent geometry, phenix.real_space_refine makes use of extra information such as secondary-structure and rotamer-specific restraints, as well as restraints or constraints on internal molecular symmetry. The re-refinement of 385 cryo-EM-derived models available in the Protein Data Bank at resolutions of 6 Å or better shows significant improvement of the models and of the fit of these models to the target maps. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29728.map.gz emd_29728.map.gz | 88.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29728-v30.xml emd-29728-v30.xml emd-29728.xml emd-29728.xml | 23.7 KB 23.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29728_fsc.xml emd_29728_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_29728.png emd_29728.png | 148.2 KB | ||
| Masks |  emd_29728_msk_1.map emd_29728_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-29728.cif.gz emd-29728.cif.gz | 7.9 KB | ||
| Others |  emd_29728_half_map_1.map.gz emd_29728_half_map_1.map.gz emd_29728_half_map_2.map.gz emd_29728_half_map_2.map.gz | 165.1 MB 165.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29728 http://ftp.pdbj.org/pub/emdb/structures/EMD-29728 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29728 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29728 | HTTPS FTP |
-Validation report
| Summary document |  emd_29728_validation.pdf.gz emd_29728_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_29728_full_validation.pdf.gz emd_29728_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_29728_validation.xml.gz emd_29728_validation.xml.gz | 20.2 KB | Display | |
| Data in CIF |  emd_29728_validation.cif.gz emd_29728_validation.cif.gz | 25.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29728 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29728 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29728 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29728 | HTTPS FTP |
-Related structure data
| Related structure data |  8g4oMC  8foiC  8g4nC  8g4xC  8g5fC  8g5gC  8g5hC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29728.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29728.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29728_msk_1.map emd_29728_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_29728_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_29728_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Native GABAA Receptor isolated from mouse brain containing two al...
| Entire | Name: Native GABAA Receptor isolated from mouse brain containing two alpha1 subunits, in complex with 8E3-Fab, didesethylflurazepam, and endogenous GABA |
|---|---|
| Components |
|
-Supramolecule #1: Native GABAA Receptor isolated from mouse brain containing two al...
| Supramolecule | Name: Native GABAA Receptor isolated from mouse brain containing two alpha1 subunits, in complex with 8E3-Fab, didesethylflurazepam, and endogenous GABA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 350 KDa |
-Macromolecule #1: Gamma-aminobutyric acid receptor subunit alpha-1
| Macromolecule | Name: Gamma-aminobutyric acid receptor subunit alpha-1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 51.818516 KDa |
| Sequence | String: MKKSRGLSDY LWAWTLILST LSGRSYGQPS QDELKDNTTV FTRILDRLLD GYDNRLRPGL GERVTEVKTD IFVTSFGPVS DHDMEYTID VFFRQSWKDE RLKFKGPMTV LRLNNLMASK IWTPDTFFHN GKKSVAHNMT MPNKLLRITE DGTLLYTMRL T VRAECPMH ...String: MKKSRGLSDY LWAWTLILST LSGRSYGQPS QDELKDNTTV FTRILDRLLD GYDNRLRPGL GERVTEVKTD IFVTSFGPVS DHDMEYTID VFFRQSWKDE RLKFKGPMTV LRLNNLMASK IWTPDTFFHN GKKSVAHNMT MPNKLLRITE DGTLLYTMRL T VRAECPMH LEDFPMDAHA CPLKFGSYAY TRAEVVYEWT REPARSVVVA EDGSRLNQYD LLGQTVDSGI VQSSTGEYVV MT THFHLKR KIGYFVIQTY LPCIMTVILS QVSFWLNRES VPARTVFGVT TVLTMTTLSI SARNSLPKVA YATAMDWFIA VCY AFVFSA LIEFATVNYF TKRGYAWDGK SVVPEKPKKV KDPLIKKNNT YAPTATSYTP NLARGDPGLA TIAKSATIEP KEVK PETKP PEPKKTFNSV SKIDRLSRIA FPLLFGIFNL VYWATYLNRE PQLKAPTPHQ UniProtKB: Gamma-aminobutyric acid receptor subunit alpha-1 |
-Macromolecule #2: Gamma-aminobutyric acid receptor subunit beta-2
| Macromolecule | Name: Gamma-aminobutyric acid receptor subunit beta-2 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 59.265312 KDa |
| Sequence | String: MWRVRKRGYF GIWSFPLIIA AVCAQSVNDP SNMSLVKETV DRLLKGYDIR LRPDFGGPPV AVGMNIDIAS IDMVSEVNMD YTLTMYFQQ AWRDKRLSYN VIPLNLTLDN RVADQLWVPD TYFLNDKKSF VHGVTVKNRM IRLHPDGTVL YGLRITTTAA C MMDLRRYP ...String: MWRVRKRGYF GIWSFPLIIA AVCAQSVNDP SNMSLVKETV DRLLKGYDIR LRPDFGGPPV AVGMNIDIAS IDMVSEVNMD YTLTMYFQQ AWRDKRLSYN VIPLNLTLDN RVADQLWVPD TYFLNDKKSF VHGVTVKNRM IRLHPDGTVL YGLRITTTAA C MMDLRRYP LDEQNCTLEI ESYGYTTDDI EFYWRGDDNA VTGVTKIELP QFSIVDYKLI TKKVVFSTGS YPRLSLSFKL KR NIGYFIL QTYMPSILIT ILSWVSFWIN YDASAARVAL GITTVLTMTT INTHLRETLP KIPYVKAIDM YLMGCFVFVF MAL LEYALV NYIFFGRGPQ RQKKAAEKAA NANNEKMRLD VNKMFYKDIK QNGTQYRSLW DPTGDLSPTR RTTNYDFSLY TMDP HENIL LSTLEIKNEM ATSEAVMGLG DPRSTMLAYD ASSIQYRKAG LPRHSFGRNA LERHVAQKKS RLRRRASQLK ITIPD LTDV NAIDRWSRIF FPVVFSFFNI VYWLYYVN UniProtKB: Gamma-aminobutyric acid receptor subunit beta-2 |
-Macromolecule #3: Gamma-aminobutyric acid receptor subunit gamma-2
| Macromolecule | Name: Gamma-aminobutyric acid receptor subunit gamma-2 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 55.160281 KDa |
| Sequence | String: MSSPNTWSIG SSVYSPVFSQ KMTLWILLLL SLYPGFTSQK SDDDYEDYAS NKTWVLTPKV PEGDVTVILN NLLEGYDNKL RPDIGVKPT LIHTDMYVNS IGPVNAINME YTIDIFFAQT WYDRRLKFNS TIKVLRLNSN MVGKIWIPDT FFRNSKKADA H WITTPNRM ...String: MSSPNTWSIG SSVYSPVFSQ KMTLWILLLL SLYPGFTSQK SDDDYEDYAS NKTWVLTPKV PEGDVTVILN NLLEGYDNKL RPDIGVKPT LIHTDMYVNS IGPVNAINME YTIDIFFAQT WYDRRLKFNS TIKVLRLNSN MVGKIWIPDT FFRNSKKADA H WITTPNRM LRIWNDGRVL YTLRLTIDAE CQLQLHNFPM DEHSCPLEFS SYGYPREEIV YQWKRSSVEV GDTRSWRLYQ FS FVGLRNT TEVVKTTSGD YVVMSVYFDL SRRMGYFTIQ TYIPCTLIVV LSWVSFWINK DAVPARTSLG ITTVLTMTTL STI ARKSLP KVSYVTAMDL FVSVCFIFVF SALVEYGTLH YFVSNRKPSK DKDKKKKNPL LRMFSFKAPT IDIRPRSATI QMNN ATHLQ ERDEEYGYEC LDGKDCASFF CCFEDCRTGA WRHGRIHIRI AKMDSYARIF FPTAFCLFNL VYWVSYLYL UniProtKB: UNIPROTKB: P22723 |
-Macromolecule #4: Heavy Chain of 8E3 Fab
| Macromolecule | Name: Heavy Chain of 8E3 Fab / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.159895 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EIQLQQSGPE LVKPGTSVKV SCKASGYSFT DYNMYWVKQS HGKSLEWIGY IDPYNADTTY NREFKGKATL TVDKSSSTAF MHLNSLTSE DSAVYYCARK RNNFYFDYWG QGTPLTVSSA KTTPPSVYPL APGCGDTTGS SVTLGCLVKG YFPESVTVTW N SGSLSSSV ...String: EIQLQQSGPE LVKPGTSVKV SCKASGYSFT DYNMYWVKQS HGKSLEWIGY IDPYNADTTY NREFKGKATL TVDKSSSTAF MHLNSLTSE DSAVYYCARK RNNFYFDYWG QGTPLTVSSA KTTPPSVYPL APGCGDTTGS SVTLGCLVKG YFPESVTVTW N SGSLSSSV HTFPALLQSG LYTMSSSVTV PSSTWPSQTV TCSVAHPASS TTVDKKSAAL EVLFQ |
-Macromolecule #5: Light Chain of 8E3 Fab
| Macromolecule | Name: Light Chain of 8E3 Fab / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.62408 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: YIVMTQSPKS MSMSLGERVT LSCRASEYVG SYVSWYQQKP EQSPKLLIYG ASNRYTGVPD RFAGSGSATD FTLTITSVQA EDLADYHCG QTYNYPTFGG GTKLEIKRAD AAPTVSIFPP SSEQLTSGGA SVVCFLNNFY PKDINVKWKI DGSERQNGVL N SWTDQDSK ...String: YIVMTQSPKS MSMSLGERVT LSCRASEYVG SYVSWYQQKP EQSPKLLIYG ASNRYTGVPD RFAGSGSATD FTLTITSVQA EDLADYHCG QTYNYPTFGG GTKLEIKRAD AAPTVSIFPP SSEQLTSGGA SVVCFLNNFY PKDINVKWKI DGSERQNGVL N SWTDQDSK DSTYSMSSTL TLTKDEYERH NSYTCEATHK TSTSPIVKSF NRNEC |
-Macromolecule #9: GAMMA-AMINO-BUTANOIC ACID
| Macromolecule | Name: GAMMA-AMINO-BUTANOIC ACID / type: ligand / ID: 9 / Number of copies: 2 / Formula: ABU |
|---|---|
| Molecular weight | Theoretical: 103.12 Da |
| Chemical component information |  ChemComp-ABU: |
-Macromolecule #10: (5M)-1-(2-aminoethyl)-7-chloro-5-(2-fluorophenyl)-1,3-dihydro-2H-...
| Macromolecule | Name: (5M)-1-(2-aminoethyl)-7-chloro-5-(2-fluorophenyl)-1,3-dihydro-2H-1,4-benzodiazepin-2-one type: ligand / ID: 10 / Number of copies: 1 / Formula: YNL |
|---|---|
| Molecular weight | Theoretical: 331.772 Da |
| Chemical component information | 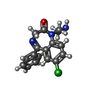 ChemComp-YNL: |
-Macromolecule #11: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 11 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 6 Component:
| ||||||||||||
| Grid | Model: Quantifoil R2/1 / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | This sample was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8g4o: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)