+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoET of HIV-2 Immature Particles | ||||||||||||||||||||||||
 Map data Map data | Odd half map | ||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | Retrovirus / Lentivirus / Gag / Capsid / HIV-2 / lattice / VIRUS LIKE PARTICLE | ||||||||||||||||||||||||
| Biological species |  Human immunodeficiency virus 2 / Human immunodeficiency virus 2 /  Human immunodeficiency virus type 2 (ISOLATE ROD) Human immunodeficiency virus type 2 (ISOLATE ROD) | ||||||||||||||||||||||||
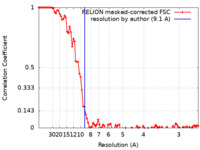
| Method | electron tomography / cryo EM / Resolution: 9.1 Å | ||||||||||||||||||||||||
 Authors Authors | Talledge N / Zhang W / Mansky ML | ||||||||||||||||||||||||
| Funding support |  United States, 7 items United States, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2023 Journal: J Mol Biol / Year: 2023Title: HIV-2 Immature Particle Morphology Provides Insights into Gag Lattice Stability and Virus Maturation. Authors: Nathaniel Talledge / Huixin Yang / Ke Shi / Raffaele Coray / Guichuan Yu / William G Arndt / Shuyu Meng / Gloria C Baxter / Luiza M Mendonça / Daniel Castaño-Díez / Hideki Aihara / Louis ...Authors: Nathaniel Talledge / Huixin Yang / Ke Shi / Raffaele Coray / Guichuan Yu / William G Arndt / Shuyu Meng / Gloria C Baxter / Luiza M Mendonça / Daniel Castaño-Díez / Hideki Aihara / Louis M Mansky / Wei Zhang /    Abstract: Retrovirus immature particle morphology consists of a membrane enclosed, pleomorphic, spherical and incomplete lattice of Gag hexamers. Previously, we demonstrated that human immunodeficiency virus ...Retrovirus immature particle morphology consists of a membrane enclosed, pleomorphic, spherical and incomplete lattice of Gag hexamers. Previously, we demonstrated that human immunodeficiency virus type 2 (HIV-2) immature particles possess a distinct and extensive Gag lattice morphology. To better understand the nature of the continuously curved hexagonal Gag lattice, we have used the single particle cryo-electron microscopy method to determine the HIV-2 Gag lattice structure for immature virions. The reconstruction map at 5.5 Å resolution revealed a stable, wineglass-shaped Gag hexamer structure with structural features consistent with other lentiviral immature Gag lattice structures. Cryo-electron tomography provided evidence for nearly complete ordered Gag lattice structures in HIV-2 immature particles. We also solved a 1.98 Å resolution crystal structure of the carboxyl-terminal domain (CTD) of the HIV-2 capsid (CA) protein that identified a structured helix 12 supported via an interaction of helix 10 in the absence of the SP1 region of Gag. Residues at the helix 10-12 interface proved critical in maintaining HIV-2 particle release and infectivity. Taken together, our findings provide the first 3D organization of HIV-2 immature Gag lattice and important insights into both HIV Gag lattice stabilization and virus maturation. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29606.map.gz emd_29606.map.gz | 29.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29606-v30.xml emd-29606-v30.xml emd-29606.xml emd-29606.xml | 19.6 KB 19.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29606_fsc.xml emd_29606_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_29606.png emd_29606.png | 59.8 KB | ||
| Masks |  emd_29606_msk_1.map emd_29606_msk_1.map | 64 MB |  Mask map Mask map | |
| Others |  emd_29606_half_map_1.map.gz emd_29606_half_map_1.map.gz emd_29606_half_map_2.map.gz emd_29606_half_map_2.map.gz | 48.8 MB 48.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29606 http://ftp.pdbj.org/pub/emdb/structures/EMD-29606 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29606 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29606 | HTTPS FTP |
-Validation report
| Summary document |  emd_29606_validation.pdf.gz emd_29606_validation.pdf.gz | 441.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_29606_full_validation.pdf.gz emd_29606_full_validation.pdf.gz | 440.8 KB | Display | |
| Data in XML |  emd_29606_validation.xml.gz emd_29606_validation.xml.gz | 15.7 KB | Display | |
| Data in CIF |  emd_29606_validation.cif.gz emd_29606_validation.cif.gz | 20.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29606 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29606 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29606 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29606 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_29606.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29606.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Odd half map | ||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||
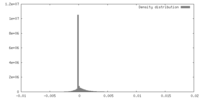
| Density |
| ||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29606_msk_1.map emd_29606_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
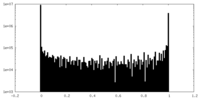
| Density Histograms |
-Half map: Even half map
| File | emd_29606_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Even half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
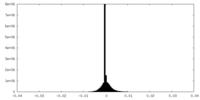
| Density Histograms |
-Half map: Odd half map
| File | emd_29606_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Odd half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
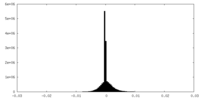
| Density Histograms |
- Sample components
Sample components
-Entire : Human immunodeficiency virus type 2 (ISOLATE ROD)
| Entire | Name:  Human immunodeficiency virus type 2 (ISOLATE ROD) Human immunodeficiency virus type 2 (ISOLATE ROD) |
|---|---|
| Components |
|
-Supramolecule #1: Human immunodeficiency virus type 2 (ISOLATE ROD)
| Supramolecule | Name: Human immunodeficiency virus type 2 (ISOLATE ROD) / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all Details: HIV-2 VLPs generated by Gag co-overexpressed with HIV-2 Env in Hek293T cells and purified from media supernatant NCBI-ID: 11720 Sci species name: Human immunodeficiency virus type 2 (ISOLATE ROD) Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Virus shell | Shell ID: 1 / Name: Capsid / Diameter: 2100.0 Å |
-Macromolecule #1: HIV-2 Immature Gag
| Macromolecule | Name: HIV-2 Immature Gag / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human immunodeficiency virus 2 / Strain: Rod Human immunodeficiency virus 2 / Strain: Rod |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGARNSVLRG KKADELERIR LRPGGKKKYR LKHIVWAANK LDRFGLAESL LESKEGCQKI LTVLDPMVP TGSENLKSLF NTVCVIWCIH AEEKVKDTEG AKQIVRRHLV AETGTAEKMP S TSRPTAPS SEKGGNYPVQ HVGGNYTHIP LSPRTLNAWV KLVEEKKFGA ...String: MGARNSVLRG KKADELERIR LRPGGKKKYR LKHIVWAANK LDRFGLAESL LESKEGCQKI LTVLDPMVP TGSENLKSLF NTVCVIWCIH AEEKVKDTEG AKQIVRRHLV AETGTAEKMP S TSRPTAPS SEKGGNYPVQ HVGGNYTHIP LSPRTLNAWV KLVEEKKFGA EVVPGFQALS EG CTPYDIN QMLNCVGDHQ AAMQIIREII NEEAAEWDVQ HPIPGPLPAG QLREPRGSDI AGT TSTVEE QIQWMFRPQN PVPVGNIYRR WIQIGLQKCV RMYNPTNILD IKQGPKEPFQ SYVD RFYKS LRAEQTDPAV KNWMTQTLLV QNANPDCKLV LKGLGMNPTL EEMLTACQGV GGPGQ KARL MAEALKEVIG PAPIPFAAAQ QRKAFKCWNC GKEGHSARQC RAPRRQGCWK CGKPGH IMT NCPDRQAGFL GLGPWGKKPR NFPVAQVPQG LTPTAPPVDP AVDLLEKYMQ QGKRQRE QR ERPYKEVTED LLHLEQGETP YREPPTEDLL HLNSLFGKDQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: 100 mM NaCl, 10 mM Tris-HCL pH 8.0, 1mM EDTA | ||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.001 kPa | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Chamber temperature: 292.15 K / Instrument: LEICA EM GP / Details: Leica GP2 Grid Plunger.. | ||||||||||||
| Sectioning | Other: NO SECTIONING | ||||||||||||
| Fiducial marker | Manufacturer: Ted pella / Diameter: 10 nm |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-8 / Number grids imaged: 2 / Number real images: 63 / Average exposure time: 2.0 sec. / Average electron dose: 1.1 e/Å2 Details: Two second exposures with a 0.25 frames/sec rate per tilt with a 1.1 e-/Angstrom^2 dose per tilt with a total tilt series dose of 45 e-/Angstroms^2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 3.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)