+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
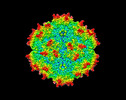
| Title | Cryo-EM Structure of empty AAV2 capsid | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | assembly / symmetry / AAV2 / capsid / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationpermeabilization of host organelle membrane involved in viral entry into host cell / symbiont entry into host cell via permeabilization of inner membrane / host cell nucleolus / T=1 icosahedral viral capsid / clathrin-dependent endocytosis of virus by host cell / virion attachment to host cell / structural molecule activity Similarity search - Function | |||||||||
| Biological species |  adeno-associated virus 2 adeno-associated virus 2 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.94 Å | |||||||||
 Authors Authors | Bennett AD / McKenna R | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: to be published Journal: to be publishedTitle: Cryo-EM Structure of genome containing AAV2 Authors: Bennett AD / Mckenna R | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29600.map.gz emd_29600.map.gz | 116.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29600-v30.xml emd-29600-v30.xml emd-29600.xml emd-29600.xml | 15.2 KB 15.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_29600.png emd_29600.png | 104.1 KB | ||
| Filedesc metadata |  emd-29600.cif.gz emd-29600.cif.gz | 6 KB | ||
| Others |  emd_29600_half_map_1.map.gz emd_29600_half_map_1.map.gz emd_29600_half_map_2.map.gz emd_29600_half_map_2.map.gz | 49.7 MB 49.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29600 http://ftp.pdbj.org/pub/emdb/structures/EMD-29600 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29600 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29600 | HTTPS FTP |
-Validation report
| Summary document |  emd_29600_validation.pdf.gz emd_29600_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_29600_full_validation.pdf.gz emd_29600_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_29600_validation.xml.gz emd_29600_validation.xml.gz | 14.5 KB | Display | |
| Data in CIF |  emd_29600_validation.cif.gz emd_29600_validation.cif.gz | 17 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29600 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29600 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29600 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29600 | HTTPS FTP |
-Related structure data
| Related structure data |  8fz0MC  8fywC  8fznC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29600.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29600.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.099 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_29600_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
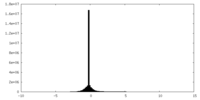
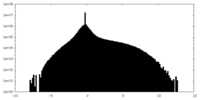
| Density Histograms |
-Half map: #2
| File | emd_29600_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
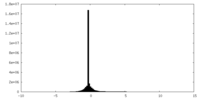
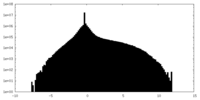
| Density Histograms |
- Sample components
Sample components
-Entire : adeno-associated virus 2
| Entire | Name:  adeno-associated virus 2 adeno-associated virus 2 |
|---|---|
| Components |
|
-Supramolecule #1: adeno-associated virus 2
| Supramolecule | Name: adeno-associated virus 2 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 10804 / Sci species name: adeno-associated virus 2 / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 3.9 MDa |
| Virus shell | Shell ID: 1 / Name: capsid / Diameter: 26.0 Å / T number (triangulation number): 1 |
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 60 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  adeno-associated virus 2 adeno-associated virus 2 |
| Molecular weight | Theoretical: 82.031352 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAADGYLPDW LEDTLSEGIR QWWKLKPGPP PPKPAERHKD DSRGLVLPGY KYLGPFNGLD KGEPVNEADA AALEHDKAYD RQLDSGDNP YLKYNHADAE FQERLKEDTS FGGNLGRAVF QAKKRVLEPL GLVEEPVKTA PGKKRPVEHS PVEPDSSSGT G KAGQQPAR ...String: MAADGYLPDW LEDTLSEGIR QWWKLKPGPP PPKPAERHKD DSRGLVLPGY KYLGPFNGLD KGEPVNEADA AALEHDKAYD RQLDSGDNP YLKYNHADAE FQERLKEDTS FGGNLGRAVF QAKKRVLEPL GLVEEPVKTA PGKKRPVEHS PVEPDSSSGT G KAGQQPAR KRLNFGQTGD ADSVPDPQPL GQPPAAPSGL GTNTMATGSG APMADNNEGA DGVGNSSGNW HCDSTWMGDR VI TTSTRTW ALPTYNNHLY KQISSQSGAS NDNHYFGYST PWGYFDFNRF HCHFSPRDWQ RLINNNWGFR PKRLNFKLFN IQV KEVTQN DGTTTIANNL TSTVQVFTDS EYQLPYVLGS AHQGCLPPFP ADVFMVPQYG YLTLNNGSQA VGRSSFYCLE YFPS QMLRT GNNFTFSYTF EDVPFHSSYA HSQSLDRLMN PLIDQYLYYL SRTNTPSGTT TQSRLQFSQA GASDIRDQSR NWLPG PCYR QQRVSKTSAD NNNSEYSWTG ATKYHLNGRD SLVNPGPAMA SHKDDEEKFF PQSGVLIFGK QGSEKTNVDI EKVMIT DEE EIRTTNPVAT EQYGSVSTNL QRGNRQAATA DVNTQGVLPG MVWQDRDVYL QGPIWAKIPH TDGHFHPSPL MGGFGLK HP PPQILIKNTP VPANPSTTFS AAKFASFITQ YSTGQVSVEI EWELQKENSK RWNPEIQYTS NYNKSVNVDF TVDTNGVY S EPRPIGTRYL TRNL UniProtKB: Capsid protein VP1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.3 / Component - Concentration: 1.0 mM / Component - Formula: PBS-MK / Component - Name: TD / Details: 1XPBS, 1 mM MgCl2, 2.5 mM KCl pH7.3 |
| Grid | Model: Quantifoil / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: DIRECT ELECTRON DE-64 (8k x 8k) / Number real images: 1674 / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.94 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cisTEM / Number images used: 6782 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8fz0: |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X