[English] 日本語
 Yorodumi
Yorodumi- EMDB-27755: Molecular Mechanism of Sialic Acid Transport Mediated by Sialin -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Molecular Mechanism of Sialic Acid Transport Mediated by Sialin | ||||||||||||
 Map data Map data | human Sialin | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | sialic acid transport / solute carrier transporter / MEMBRANE PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsialic acid:proton symporter activity / D-glucuronate transmembrane transporter activity / Defective SLC17A5 causes Salla disease (SD) and ISSD / sialic acid transport / Organic anion transporters / sialic acid transmembrane transporter activity / carbohydrate:proton symporter activity / Sialic acid metabolism / monoatomic anion transport / amino acid transport ...sialic acid:proton symporter activity / D-glucuronate transmembrane transporter activity / Defective SLC17A5 causes Salla disease (SD) and ISSD / sialic acid transport / Organic anion transporters / sialic acid transmembrane transporter activity / carbohydrate:proton symporter activity / Sialic acid metabolism / monoatomic anion transport / amino acid transport / monoatomic ion transport / response to bacterium / synaptic vesicle membrane / basolateral plasma membrane / lysosome / lysosomal membrane / membrane / plasma membrane / cytosol Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | ||||||||||||
 Authors Authors | Hu W / Zheng H | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2023 Journal: Sci Adv / Year: 2023Title: The molecular mechanism of sialic acid transport mediated by Sialin. Authors: Wenxin Hu / Congwu Chi / Kunhua Song / Hongjin Zheng /  Abstract: Malfunction of the sialic acid transporter caused by various genetic mutations in the gene encoding Sialin leads to a spectrum of neurodegenerative conditions called free sialic acid storage ...Malfunction of the sialic acid transporter caused by various genetic mutations in the gene encoding Sialin leads to a spectrum of neurodegenerative conditions called free sialic acid storage disorders. Unfortunately, how Sialin transports sialic acid/proton (H) and how pathogenic mutations impair its function are poorly defined. Here, we present the structure of human Sialin in an inward-facing partially open conformation determined by cryo-electron microscopy, representing the first high-resolution structure of any human SLC17 member. Our analysis reveals two unique features in Sialin: (i) The H coupling/sensing requires two highly conserved Glu residues (E171 and E175) instead of one (E175) as implied in previous studies; and (ii) the normal function of Sialin requires the stabilization of a cytosolic helix, which has not been noticed in the literature. By mapping known pathogenic mutations, we provide mechanistic explanations for corresponding functional defects. We propose a structure-based mechanism for sialic acid transport mediated by Sialin. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27755.map.gz emd_27755.map.gz | 118 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27755-v30.xml emd-27755-v30.xml emd-27755.xml emd-27755.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27755.png emd_27755.png | 61.6 KB | ||
| Filedesc metadata |  emd-27755.cif.gz emd-27755.cif.gz | 5.4 KB | ||
| Others |  emd_27755_half_map_1.map.gz emd_27755_half_map_1.map.gz emd_27755_half_map_2.map.gz emd_27755_half_map_2.map.gz | 115.9 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27755 http://ftp.pdbj.org/pub/emdb/structures/EMD-27755 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27755 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27755 | HTTPS FTP |
-Validation report
| Summary document |  emd_27755_validation.pdf.gz emd_27755_validation.pdf.gz | 880.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27755_full_validation.pdf.gz emd_27755_full_validation.pdf.gz | 880.5 KB | Display | |
| Data in XML |  emd_27755_validation.xml.gz emd_27755_validation.xml.gz | 13.8 KB | Display | |
| Data in CIF |  emd_27755_validation.cif.gz emd_27755_validation.cif.gz | 16.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27755 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27755 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27755 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27755 | HTTPS FTP |
-Related structure data
| Related structure data |  8dwiMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27755.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27755.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | human Sialin | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
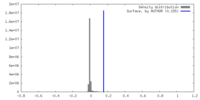
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map B
| File | emd_27755_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
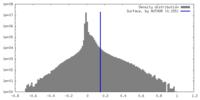
| Density Histograms |
-Half map: half map A
| File | emd_27755_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
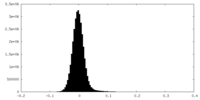
| Density Histograms |
- Sample components
Sample components
-Entire : Sialin complex with Fab 8B1
| Entire | Name: Sialin complex with Fab 8B1 |
|---|---|
| Components |
|
-Supramolecule #1: Sialin complex with Fab 8B1
| Supramolecule | Name: Sialin complex with Fab 8B1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Sialin
| Macromolecule | Name: Sialin / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 54.684168 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRSPVRDLAR NDGEESTDRT PLLPGAPRAE AAPVCCSARY NLAILAFFGF FIVYALRVNL SVALVDMVDS NTTLEDNRTS KACPEHSAP IKVHHNQTGK KYQWDAETQG WILGSFFYGY IITQIPGGYV ASKIGGKMLL GFGILGTAVL TLFTPIAADL G VGPLIVLR ...String: MRSPVRDLAR NDGEESTDRT PLLPGAPRAE AAPVCCSARY NLAILAFFGF FIVYALRVNL SVALVDMVDS NTTLEDNRTS KACPEHSAP IKVHHNQTGK KYQWDAETQG WILGSFFYGY IITQIPGGYV ASKIGGKMLL GFGILGTAVL TLFTPIAADL G VGPLIVLR ALEGLGEGVT FPAMHAMWSS WAPPLERSKL LSISYAGAQL GTVISLPLSG IICYYMNWTY VFYFFGTIGI FW FLLWIWL VSDTPQKHKR ISHYEKEYIL SSLRNQLSSQ KSVPWVPILK SLPLWAIVVA HFSYNWTFYT LLTLLPTYMK EIL RFNVQE NGFLSSLPYL GSWLCMILSG QAADNLRAKW NFSTLCVRRI FSLIGMIGPA VFLVAAGFIG CDYSLAVAFL TIST TLGGF CSSGFSINHL DIAPSYAGIL LGITNTFATI PGMVGPVIAK SLTPDNTVGE WQTVFYIAAA INVFGAIFFT LFAKG EVQN WALNDHHGHR H UniProtKB: Sialin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 57.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.4 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 394078 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)