+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2763 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of a partial yeast 48S preinitiation complex | |||||||||
 Map data Map data | Reconstruction of a partial 48S preinitiation complex from yeast. In order to see all the components it is recommended to apply a gaussian filter of 1.5 width and then visualize it at a contour level of 0.02. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Eukaryotic translation initiation / 48S / small ribosome subunit. | |||||||||
| Function / homology |  Function and homology information Function and homology informationformation of translation initiation ternary complex / Recycling of eIF2:GDP / Cellular response to mitochondrial stress / ABC-family proteins mediated transport / methionyl-initiator methionine tRNA binding / translation reinitiation / eukaryotic translation initiation factor 2 complex / formation of cytoplasmic translation initiation complex / multi-eIF complex / protein-synthesizing GTPase ...formation of translation initiation ternary complex / Recycling of eIF2:GDP / Cellular response to mitochondrial stress / ABC-family proteins mediated transport / methionyl-initiator methionine tRNA binding / translation reinitiation / eukaryotic translation initiation factor 2 complex / formation of cytoplasmic translation initiation complex / multi-eIF complex / protein-synthesizing GTPase / eukaryotic 43S preinitiation complex / formation of translation preinitiation complex / eukaryotic 48S preinitiation complex / positive regulation of translational fidelity / Formation of the ternary complex, and subsequently, the 43S complex / Translation initiation complex formation / Ribosomal scanning and start codon recognition / Formation of a pool of free 40S subunits / L13a-mediated translational silencing of Ceruloplasmin expression / ribosomal small subunit binding / 90S preribosome / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / regulation of translational fidelity / translation regulator activity / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / translation initiation factor binding / translation initiation factor activity / rescue of stalled ribosome / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA / small-subunit processome / translational initiation / protein kinase C binding / positive regulation of apoptotic signaling pathway / maintenance of translational fidelity / cytoplasmic stress granule / modification-dependent protein catabolic process / rRNA processing / protein tag activity / double-stranded RNA binding / ribosomal small subunit biogenesis / small ribosomal subunit rRNA binding / ribosome binding / ribosomal small subunit assembly / small ribosomal subunit / cytosolic small ribosomal subunit / cytoplasmic translation / cytosolic large ribosomal subunit / rRNA binding / ribosome / protein ubiquitination / structural constituent of ribosome / ribonucleoprotein complex / positive regulation of protein phosphorylation / translation / GTPase activity / mRNA binding / ubiquitin protein ligase binding / GTP binding / nucleolus / protein kinase binding / RNA binding / zinc ion binding / nucleus / metal ion binding / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Kluyveromyces lactis (yeast) / Kluyveromyces lactis (yeast) /  | |||||||||
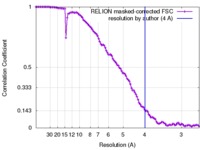
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Hussain T / Llacer JL / Fernandez IS / Savva CG / Ramakrishnan V | |||||||||
 Citation Citation |  Journal: Cell / Year: 2014 Journal: Cell / Year: 2014Title: Structural changes enable start codon recognition by the eukaryotic translation initiation complex. Authors: Tanweer Hussain / Jose L Llácer / Israel S Fernández / Antonio Munoz / Pilar Martin-Marcos / Christos G Savva / Jon R Lorsch / Alan G Hinnebusch / V Ramakrishnan /   Abstract: During eukaryotic translation initiation, initiator tRNA does not insert fully into the P decoding site on the 40S ribosomal subunit. This conformation (POUT) is compatible with scanning mRNA for the ...During eukaryotic translation initiation, initiator tRNA does not insert fully into the P decoding site on the 40S ribosomal subunit. This conformation (POUT) is compatible with scanning mRNA for the AUG start codon. Base pairing with AUG is thought to promote isomerization to a more stable conformation (PIN) that arrests scanning and promotes dissociation of eIF1 from the 40S subunit. Here, we present a cryoEM reconstruction of a yeast preinitiation complex at 4.0 Å resolution with initiator tRNA in the PIN state, prior to eIF1 release. The structure reveals stabilization of the codon-anticodon duplex by the N-terminal tail of eIF1A, changes in the structure of eIF1 likely instrumental in its subsequent release, and changes in the conformation of eIF2. The mRNA traverses the entire mRNA cleft and makes connections to the regulatory domain of eIF2?, eIF1A, and ribosomal elements that allow recognition of context nucleotides surrounding the AUG codon. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2763.map.gz emd_2763.map.gz | 96.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2763-v30.xml emd-2763-v30.xml emd-2763.xml emd-2763.xml | 15.9 KB 15.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_2763_fsc.xml emd_2763_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  EMD-2763.png EMD-2763.png emd_2763.png emd_2763.png | 151.3 KB 151.3 KB | ||
| Others |  emd_2763_additional_1.map.gz emd_2763_additional_1.map.gz | 96.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2763 http://ftp.pdbj.org/pub/emdb/structures/EMD-2763 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2763 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2763 | HTTPS FTP |
-Validation report
| Summary document |  emd_2763_validation.pdf.gz emd_2763_validation.pdf.gz | 430.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2763_full_validation.pdf.gz emd_2763_full_validation.pdf.gz | 429.7 KB | Display | |
| Data in XML |  emd_2763_validation.xml.gz emd_2763_validation.xml.gz | 11.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2763 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2763 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2763 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2763 | HTTPS FTP |
-Related structure data
| Related structure data |  3j81MC  2764C  3j80C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_2763.map.gz / Format: CCP4 / Size: 100.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2763.map.gz / Format: CCP4 / Size: 100.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of a partial 48S preinitiation complex from yeast. In order to see all the components it is recommended to apply a gaussian filter of 1.5 width and then visualize it at a contour level of 0.02. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Supplemental map: emd 2763 additional 1.map
| File | emd_2763_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Partial yeast 48S preinitiation complex
| Entire | Name: Partial yeast 48S preinitiation complex |
|---|---|
| Components |
|
-Supramolecule #1000: Partial yeast 48S preinitiation complex
| Supramolecule | Name: Partial yeast 48S preinitiation complex / type: sample / ID: 1000 / Oligomeric state: 1 / Number unique components: 6 |
|---|---|
| Molecular weight | Theoretical: 1.4 MDa |
-Supramolecule #1: Ribosome small subunit
| Supramolecule | Name: Ribosome small subunit / type: complex / ID: 1 / Name.synonym: 40S / Recombinant expression: No / Ribosome-details: ribosome-eukaryote: SSU 40S, SSU RNA 18S |
|---|---|
| Source (natural) | Organism:  Kluyveromyces lactis (yeast) Kluyveromyces lactis (yeast) |
| Molecular weight | Theoretical: 1.2 MDa |
-Macromolecule #1: Eukaryotic initiation factor 1
| Macromolecule | Name: Eukaryotic initiation factor 1 / type: protein_or_peptide / ID: 1 / Name.synonym: eIF1 / Number of copies: 1 / Oligomeric state: Monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.3 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Eukaryotic translation initiation factor eIF-1 |
-Macromolecule #2: Eukaryotic initiation factor 1A
| Macromolecule | Name: Eukaryotic initiation factor 1A / type: protein_or_peptide / ID: 2 / Name.synonym: eIF1A / Number of copies: 1 / Oligomeric state: Monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 17.4 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Eukaryotic translation initiation factor 1A |
-Macromolecule #3: Eukaryotic initiation factor 2
| Macromolecule | Name: Eukaryotic initiation factor 2 / type: protein_or_peptide / ID: 3 / Name.synonym: eIF2 Details: Uniprot codes are: alpha-P20459 beta-P09064 gamma-P32481 Number of copies: 1 / Oligomeric state: Three subunits, alpha, beta, gamma / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 124 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #4: Initiator transfer RNA
| Macromolecule | Name: Initiator transfer RNA / type: rna / ID: 4 / Name.synonym: Met-tRNAi Details: Doble mutation (G31U:C39A) when compared with yeast WT initiator tRNA. Classification: TRANSFER / Structure: SINGLE STRANDED / Synthetic?: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23 KDa |
| Sequence | String: AGCGCCGUGG CGCAGUGGAA GCGCGCAGGU CUCAUAAACC UGAUGUCCUC GGAUCGAAAC CGAGCGGCGC UACCA |
-Macromolecule #5: Messenger RNA
| Macromolecule | Name: Messenger RNA / type: rna / ID: 5 / Name.synonym: mRNA / Classification: OTHER / Structure: SINGLE STRANDED / Synthetic?: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 7.4 KDa |
| Sequence | String: GGAAUCUCUC UCUAUGCUCU CUCUC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.17 mg/mL |
|---|---|
| Buffer | pH: 6.5 Details: 20mM MES-KOH, 40mM K-acetate, 10mM NH4-acetate, 8mM Mg-acetate, 2mM DTT |
| Grid | Details: Quantifoil R2/2 400 mesh copper grids with 4-5 nm thin carbon on top |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 120 K / Instrument: FEI VITROBOT MARK I / Timed resolved state: 30 second incubation time / Method: Blot for 2.5 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 59,000 times magnification |
| Details | Complete dataset was collected in 5 non-consecutive sessions |
| Date | Oct 28, 2013 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON II (4k x 4k) / Number real images: 1791 / Average electron dose: 28 e/Å2 Details: Complete dataset was collected in 5 non-consecutive sessions |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 104478 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 0.004 µm / Nominal defocus min: 0.0016 µm / Nominal magnification: 78000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID:  3u5b Chain - Chain ID: 2 |
|---|---|
| Software | Name: Chimera, refmac |
| Refinement | Space: RECIPROCAL / Protocol: FLEXIBLE FIT / Target criteria: R-factor, FSC |
| Output model |  PDB-3j81: |
-Atomic model buiding 2
| Initial model | PDB ID:  3u5c |
|---|---|
| Software | Name: Chimera, refmac |
| Refinement | Space: RECIPROCAL / Protocol: FLEXIBLE FIT / Target criteria: R-factor, FSC |
| Output model |  PDB-3j81: |
 Movie
Movie Controller
Controller




























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)