[English] 日本語
 Yorodumi
Yorodumi- EMDB-26726: PARL-cleaved Skd3 (human ClpB) E455Q Nucleotide Binding Domain he... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | PARL-cleaved Skd3 (human ClpB) E455Q Nucleotide Binding Domain hexamer bound to ATPgammaS, closed conformation | |||||||||||||||||||||
 Map data Map data | PARL-cleaved Skd3 (human ClpB) E455Q Nucleotide Binding Domain hexamer bound to ATPgammaS in the closed conformation from 3DVA | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | AAA+ ATPase / Chaperone / Mitochondria / Protein Folding | |||||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||||||||
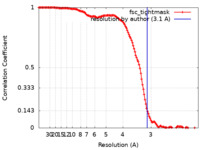
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||||||||||||||
 Authors Authors | Gupta A / Lentzsch AM / Siegel AS / Yu Z / Lu C / Chio US / Cheng Y / Shan S | |||||||||||||||||||||
| Funding support |  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2023 Journal: Sci Adv / Year: 2023Title: Dodecamer assembly of a metazoan AAA chaperone couples substrate extraction to refolding. Authors: Arpit Gupta / Alfred M Lentzsch / Alex Siegel / Zanlin Yu / Un Seng Chio / Yifan Cheng / Shu-Ou Shan /  Abstract: Ring-forming AAA chaperones solubilize protein aggregates and protect organisms from proteostatic stress. In metazoans, the AAA chaperone Skd3 in the mitochondrial intermembrane space (IMS) is ...Ring-forming AAA chaperones solubilize protein aggregates and protect organisms from proteostatic stress. In metazoans, the AAA chaperone Skd3 in the mitochondrial intermembrane space (IMS) is critical for human health and efficiently refolds aggregated proteins, but its underlying mechanism is poorly understood. Here, we show that Skd3 harbors both disaggregase and protein refolding activities enabled by distinct assembly states. High-resolution structures of Skd3 hexamers in distinct conformations capture ratchet-like motions that mediate substrate extraction. Unlike previously described disaggregases, Skd3 hexamers further assemble into dodecameric cages in which solubilized substrate proteins can attain near-native states. Skd3 mutants defective in dodecamer assembly retain disaggregase activity but are impaired in client refolding, linking the disaggregase and refolding activities to the hexameric and dodecameric states of Skd3, respectively. We suggest that Skd3 is a combined disaggregase and foldase, and this property is particularly suited to meet the complex proteostatic demands in the mitochondrial IMS. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26726.map.gz emd_26726.map.gz | 108 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26726-v30.xml emd-26726-v30.xml emd-26726.xml emd-26726.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26726_fsc.xml emd_26726_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_26726.png emd_26726.png | 68.6 KB | ||
| Filedesc metadata |  emd-26726.cif.gz emd-26726.cif.gz | 5 KB | ||
| Others |  emd_26726_half_map_1.map.gz emd_26726_half_map_1.map.gz emd_26726_half_map_2.map.gz emd_26726_half_map_2.map.gz | 200.6 MB 200.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26726 http://ftp.pdbj.org/pub/emdb/structures/EMD-26726 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26726 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26726 | HTTPS FTP |
-Validation report
| Summary document |  emd_26726_validation.pdf.gz emd_26726_validation.pdf.gz | 697.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_26726_full_validation.pdf.gz emd_26726_full_validation.pdf.gz | 697.4 KB | Display | |
| Data in XML |  emd_26726_validation.xml.gz emd_26726_validation.xml.gz | 20.5 KB | Display | |
| Data in CIF |  emd_26726_validation.cif.gz emd_26726_validation.cif.gz | 25.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26726 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26726 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26726 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26726 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_26726.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26726.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PARL-cleaved Skd3 (human ClpB) E455Q Nucleotide Binding Domain hexamer bound to ATPgammaS in the closed conformation from 3DVA | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0428 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: PARL-cleaved Skd3 (human ClpB) E455Q Nucleotide Binding Domain...
| File | emd_26726_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PARL-cleaved Skd3 (human ClpB) E455Q Nucleotide Binding Domain hexamer bound to ATPgammaS in the closed conformation, half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: PARL-cleaved Skd3 (human ClpB) E455Q Nucleotide Binding Domain...
| File | emd_26726_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PARL-cleaved Skd3 (human ClpB) E455Q Nucleotide Binding Domain hexamer bound to ATPgammaS in the closed conformation, half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PARL-cleaved Skd3
| Entire | Name: PARL-cleaved Skd3 |
|---|---|
| Components |
|
-Supramolecule #1: PARL-cleaved Skd3
| Supramolecule | Name: PARL-cleaved Skd3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 397 KDa |
-Macromolecule #1: PARL-cleaved Skd3
| Macromolecule | Name: PARL-cleaved Skd3 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: YSKSPSNKDA ALLEAARANN MQEVSRLLSE GADVNAKHRL GWTALMVAAI NRNNSVVQVL LAAGADPNLG DDFSSVYKTA KEQGIHSLED GGQDGASRHI TNQWTSALEF RRWLGLPAGV LITREDDFNN RLNNRASFKG CTALHYAVLA DDYRTVKELL DGGANPLQRN ...String: YSKSPSNKDA ALLEAARANN MQEVSRLLSE GADVNAKHRL GWTALMVAAI NRNNSVVQVL LAAGADPNLG DDFSSVYKTA KEQGIHSLED GGQDGASRHI TNQWTSALEF RRWLGLPAGV LITREDDFNN RLNNRASFKG CTALHYAVLA DDYRTVKELL DGGANPLQRN EMGHTPLDYA REGEVMKLLR TSEAKYQEKQ RKREAEERRR FPLEQRLKEH IIGQESAIAT VGAAIRRKEN GWYDEEHPLV FLFLGSSGIG KTELAKQTAK YMHKDAKKGF IRLDMSEFQE RHEVAKFIGS PPGYVGHEEG GQLTKKLKQC PNAVVLFDQV DKAHPDVLTI MLQLFDEGRL TDGKGKTIDC KDAIFIMTSN VASDEIAQHA LQLRQEALEM SRNRIAENLG DVQISDKITI SKNFKENVIR PILKAHFRRD EFLGRINEIV YFLPFCHSEL IQLVNKELNF WAKRAKQRHN ITLLWDREVA DVLVDGYNVH YGARSIKHEV ERRVVNQLAA AYEQDLLPGG CTLRITVEDS DKQLLKSPEL PSPQAEKRLP KLRLEIIDKD SKTRRLDIRA PLHPEKVCNT I |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: -2.0 µm / Nominal defocus min: -1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)