[English] 日本語
 Yorodumi
Yorodumi- EMDB-26689: Locally refined 5S rRNP map from the nucleoplasmic pre-60S interm... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Locally refined 5S rRNP map from the nucleoplasmic pre-60S intermediate of the Nog2 containing pre-rotation state | ||||||||||||
 Map data Map data | Nog2-containing nuclear pre-60S intermediate local map centered on the 5S rRNP | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | ribosome biogenesis / K-loop GTPase / GTPase / RIBOSOME / 5S rRNP | ||||||||||||
| Biological species |  | ||||||||||||
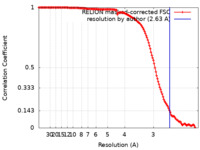
| Method | single particle reconstruction / cryo EM / Resolution: 2.63 Å | ||||||||||||
 Authors Authors | Sekulski K / Cruz VE / Weirich CS / Erzberger JP | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: rRNA methylation by Spb1 regulates the GTPase activity of Nog2 during 60S ribosomal subunit assembly. Authors: Kamil Sekulski / Victor Emmanuel Cruz / Christine S Weirich / Jan P Erzberger /  Abstract: Biogenesis of the large ribosomal (60S) subunit involves the assembly of three rRNAs and 46 proteins, a process requiring approximately 70 ribosome biogenesis factors (RBFs) that bind and release the ...Biogenesis of the large ribosomal (60S) subunit involves the assembly of three rRNAs and 46 proteins, a process requiring approximately 70 ribosome biogenesis factors (RBFs) that bind and release the pre-60S at specific steps along the assembly pathway. The methyltransferase Spb1 and the K-loop GTPase Nog2 are essential RBFs that engage the rRNA A-loop during sequential steps in 60S maturation. Spb1 methylates the A-loop nucleotide G2922 and a catalytically deficient mutant strain (spb1) has a severe 60S biogenesis defect. However, the assembly function of this modification is currently unknown. Here, we present cryo-EM reconstructions that reveal that unmethylated G2922 leads to the premature activation of Nog2 GTPase activity and capture a Nog2-GDP-AlF transition state structure that implicates the direct involvement of unmodified G2922 in Nog2 GTPase activation. Genetic suppressors and in vivo imaging indicate that premature GTP hydrolysis prevents the efficient binding of Nog2 to early nucleoplasmic 60S intermediates. We propose that G2922 methylation levels regulate Nog2 recruitment to the pre-60S near the nucleolar/nucleoplasmic phase boundary, forming a kinetic checkpoint to regulate 60S production. Our approach and findings provide a template to study the GTPase cycles and regulatory factor interactions of the other K-loop GTPases involved in ribosome assembly. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26689.map.gz emd_26689.map.gz | 223.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26689-v30.xml emd-26689-v30.xml emd-26689.xml emd-26689.xml | 26.2 KB 26.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26689_fsc.xml emd_26689_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_26689.png emd_26689.png | 68.5 KB | ||
| Masks |  emd_26689_msk_1.map emd_26689_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-26689.cif.gz emd-26689.cif.gz | 4.9 KB | ||
| Others |  emd_26689_half_map_1.map.gz emd_26689_half_map_1.map.gz emd_26689_half_map_2.map.gz emd_26689_half_map_2.map.gz | 212.9 MB 212.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26689 http://ftp.pdbj.org/pub/emdb/structures/EMD-26689 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26689 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26689 | HTTPS FTP |
-Validation report
| Summary document |  emd_26689_validation.pdf.gz emd_26689_validation.pdf.gz | 886.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_26689_full_validation.pdf.gz emd_26689_full_validation.pdf.gz | 886 KB | Display | |
| Data in XML |  emd_26689_validation.xml.gz emd_26689_validation.xml.gz | 21.6 KB | Display | |
| Data in CIF |  emd_26689_validation.cif.gz emd_26689_validation.cif.gz | 28.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26689 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26689 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26689 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26689 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_26689.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26689.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Nog2-containing nuclear pre-60S intermediate local map centered on the 5S rRNP | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_26689_msk_1.map emd_26689_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-map 1 of Nog2-containing nuclear pre-60S intermediate local...
| File | emd_26689_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 1 of Nog2-containing nuclear pre-60S intermediate local map centered on the 5S rRNP | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-map 2 of Nog2-containing nuclear pre-60S intermediate local...
| File | emd_26689_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 2 of Nog2-containing nuclear pre-60S intermediate local map centered on the 5S rRNP | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Local 5S map from the nucleolar pre-60S intermediate purified wit...
| Entire | Name: Local 5S map from the nucleolar pre-60S intermediate purified with tags on Tif6 and Nog2. |
|---|---|
| Components |
|
-Supramolecule #1: Local 5S map from the nucleolar pre-60S intermediate purified wit...
| Supramolecule | Name: Local 5S map from the nucleolar pre-60S intermediate purified with tags on Tif6 and Nog2. type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#34, #36-#60 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 0.3 / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 5580 / Average exposure time: 0.05 sec. / Average electron dose: 1.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 2.2 µm / Calibrated defocus min: 0.9 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)