[English] 日本語
 Yorodumi
Yorodumi- EMDB-25418: Post translocation, non-rotated 70S ribosome with EF-G dissociate... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25418 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Post translocation, non-rotated 70S ribosome with EF-G dissociated (Structure VII) | |||||||||||||||
 Map data Map data | map used in refinement of VII | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Ribosome / Translocation / Classical | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationstringent response / negative regulation of cytoplasmic translational initiation / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / misfolded RNA binding / Group I intron splicing / RNA folding / transcriptional attenuation / translational termination / endoribonuclease inhibitor activity ...stringent response / negative regulation of cytoplasmic translational initiation / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / misfolded RNA binding / Group I intron splicing / RNA folding / transcriptional attenuation / translational termination / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity / four-way junction DNA binding / negative regulation of cytoplasmic translation / DnaA-L2 complex / regulation of mRNA stability / translation repressor activity / negative regulation of translational initiation / negative regulation of DNA-templated DNA replication initiation / mRNA regulatory element binding translation repressor activity / positive regulation of RNA splicing / regulation of DNA-templated transcription elongation / transcription elongation factor complex / response to reactive oxygen species / cytosolic ribosome assembly / ribosome assembly / assembly of large subunit precursor of preribosome / transcription antitermination / DNA endonuclease activity / translational initiation / regulation of cell growth / DNA-templated transcription termination / response to radiation / maintenance of translational fidelity / mRNA 5'-UTR binding / regulation of translation / large ribosomal subunit / ribosomal small subunit assembly / transferase activity / ribosome biogenesis / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / response to antibiotic / negative regulation of DNA-templated transcription / hydrolase activity / mRNA binding / DNA binding / RNA binding / zinc ion binding / membrane / cytoplasm / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||||||||
 Authors Authors | Carbone CE / Korostelev AA | |||||||||||||||
| Funding support |  Czech Republic, 4 items Czech Republic, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP. Authors: Christine E Carbone / Anna B Loveland / Howard B Gamper / Ya-Ming Hou / Gabriel Demo / Andrei A Korostelev /   Abstract: During translation, a conserved GTPase elongation factor-EF-G in bacteria or eEF2 in eukaryotes-translocates tRNA and mRNA through the ribosome. EF-G has been proposed to act as a flexible motor that ...During translation, a conserved GTPase elongation factor-EF-G in bacteria or eEF2 in eukaryotes-translocates tRNA and mRNA through the ribosome. EF-G has been proposed to act as a flexible motor that propels tRNA and mRNA movement, as a rigid pawl that biases unidirectional translocation resulting from ribosome rearrangements, or by various combinations of motor- and pawl-like mechanisms. Using time-resolved cryo-EM, we visualized GTP-catalyzed translocation without inhibitors, capturing elusive structures of ribosome•EF-G intermediates at near-atomic resolution. Prior to translocation, EF-G binds near peptidyl-tRNA, while the rotated 30S subunit stabilizes the EF-G GTPase center. Reverse 30S rotation releases Pi and translocates peptidyl-tRNA and EF-G by ~20 Å. An additional 4-Å translocation initiates EF-G dissociation from a transient ribosome state with highly swiveled 30S head. The structures visualize how nearly rigid EF-G rectifies inherent and spontaneous ribosomal dynamics into tRNA-mRNA translocation, whereas GTP hydrolysis and Pi release drive EF-G dissociation. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25418.map.gz emd_25418.map.gz | 317.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25418-v30.xml emd-25418-v30.xml emd-25418.xml emd-25418.xml | 80.5 KB 80.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25418.png emd_25418.png | 193.2 KB | ||
| Filedesc metadata |  emd-25418.cif.gz emd-25418.cif.gz | 13.7 KB | ||
| Others |  emd_25418_additional_1.map.gz emd_25418_additional_1.map.gz emd_25418_additional_2.map.gz emd_25418_additional_2.map.gz emd_25418_half_map_1.map.gz emd_25418_half_map_1.map.gz emd_25418_half_map_2.map.gz emd_25418_half_map_2.map.gz | 317.9 MB 23.8 MB 143.9 MB 143.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25418 http://ftp.pdbj.org/pub/emdb/structures/EMD-25418 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25418 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25418 | HTTPS FTP |
-Related structure data
| Related structure data |  7st2MC  7ss9C  7ssdC  7sslC  7ssnC  7ssoC  7sswC  7st6C  7st7C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25418.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25418.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map used in refinement of VII | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.87 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
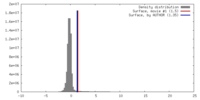
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: map of VII after focused classification (30A mask...
| File | emd_25418_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map of VII after focused classification (30A mask around the Shine-Dalgarno for 200 cycles) into 3 classes. Used for local modelling and refinement of the Shine-Dalgarno helix. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: blocfiltered version of map isolated through focused masking...
| File | emd_25418_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | blocfiltered version of map isolated through focused masking of the Shine-Dalgarno helix. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_25418_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_25418_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Post translocation, non-rotated 70S ribosome with EF-G dissociate...
+Supramolecule #1: Post translocation, non-rotated 70S ribosome with EF-G dissociate...
+Macromolecule #1: 16S rRNA
+Macromolecule #2: 23S rRNA
+Macromolecule #3: 5S rRNA
+Macromolecule #4: tRNA fMet
+Macromolecule #37: mRNA
+Macromolecule #38: tRNA Pro
+Macromolecule #5: 50S ribosomal protein L2
+Macromolecule #6: 50S ribosomal protein L3
+Macromolecule #7: 50S ribosomal protein L4
+Macromolecule #8: 50S ribosomal protein L5
+Macromolecule #9: 50S ribosomal protein L6
+Macromolecule #10: 50S ribosomal protein L9
+Macromolecule #11: 50S ribosomal protein L1
+Macromolecule #12: 50S ribosomal protein L10
+Macromolecule #13: 50S ribosomal protein L11
+Macromolecule #14: 50S ribosomal protein L13
+Macromolecule #15: 50S ribosomal protein L14
+Macromolecule #16: 50S ribosomal protein L15
+Macromolecule #17: 50S ribosomal protein L16
+Macromolecule #18: 50S ribosomal protein L17
+Macromolecule #19: 50S ribosomal protein L18
+Macromolecule #20: 50S ribosomal protein L19
+Macromolecule #21: 50S ribosomal protein L20
+Macromolecule #22: 50S ribosomal protein L21
+Macromolecule #23: 50S ribosomal protein L22
+Macromolecule #24: 50S ribosomal protein L23
+Macromolecule #25: 50S ribosomal protein L24
+Macromolecule #26: 50S ribosomal protein L25
+Macromolecule #27: 50S ribosomal protein L27
+Macromolecule #28: 50S ribosomal protein L28
+Macromolecule #29: 50S ribosomal protein L29
+Macromolecule #30: 50S ribosomal protein L30
+Macromolecule #31: 50S ribosomal protein L31
+Macromolecule #32: 50S ribosomal protein L32
+Macromolecule #33: 50S ribosomal protein L33
+Macromolecule #34: 50S ribosomal protein L34
+Macromolecule #35: 50S ribosomal protein L35
+Macromolecule #36: 50S ribosomal protein L36
+Macromolecule #39: 30S ribosomal protein S2
+Macromolecule #40: 30S ribosomal protein S3
+Macromolecule #41: 30S ribosomal protein S4
+Macromolecule #42: 30S ribosomal protein S5
+Macromolecule #43: 30S ribosomal protein S6
+Macromolecule #44: 30S ribosomal protein S7
+Macromolecule #45: 30S ribosomal protein S8
+Macromolecule #46: 30S ribosomal protein S9
+Macromolecule #47: 30S ribosomal protein S10
+Macromolecule #48: 30S ribosomal protein S11
+Macromolecule #49: 30S ribosomal protein S12
+Macromolecule #50: 30S ribosomal protein S13
+Macromolecule #51: 30S ribosomal protein S14
+Macromolecule #52: 30S ribosomal protein S15
+Macromolecule #53: 30S ribosomal protein S16
+Macromolecule #54: 30S ribosomal protein S17
+Macromolecule #55: 30S ribosomal protein S18
+Macromolecule #56: 30S ribosomal protein S19
+Macromolecule #57: 30S ribosomal protein S20
+Macromolecule #58: 30S ribosomal protein S21
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 30.4 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.9 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: FREALIGN / Number images used: 55457 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: FREALIGN |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: FREALIGN |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)