[English] 日本語
 Yorodumi
Yorodumi- EMDB-25206: Cryo-EM structure of Torpedo acetylcholine receptor in complex wi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25206 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Torpedo acetylcholine receptor in complex with carbachol, desensitized state | |||||||||
 Map data Map data | Cryo-EM structure of Torpedo acetylcholine receptor in complex with carbachol, desensitized state | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationacetylcholine-gated monoatomic cation-selective channel activity / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / transmembrane signaling receptor activity / postsynaptic membrane / neuron projection Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.77 Å | |||||||||
 Authors Authors | Rahman MM / Basta T / Teng J / Lee M / Worrell BT / Stowell MHB / Hibbs RE | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2022 Journal: Nat Struct Mol Biol / Year: 2022Title: Structural mechanism of muscle nicotinic receptor desensitization and block by curare. Authors: Md Mahfuzur Rahman / Tamara Basta / Jinfeng Teng / Myeongseon Lee / Brady T Worrell / Michael H B Stowell / Ryan E Hibbs /  Abstract: Binding of the neurotransmitter acetylcholine to its receptors on muscle fibers depolarizes the membrane and thereby triggers muscle contraction. We sought to understand at the level of three- ...Binding of the neurotransmitter acetylcholine to its receptors on muscle fibers depolarizes the membrane and thereby triggers muscle contraction. We sought to understand at the level of three-dimensional structure how agonists and antagonists alter nicotinic acetylcholine receptor conformation. We used the muscle-type receptor from the Torpedo ray to first define the structure of the receptor in a resting, activatable state. We then determined the receptor structure bound to the agonist carbachol, which stabilizes an asymmetric, closed channel desensitized state. We find conformational changes in a peripheral membrane helix are tied to recovery from desensitization. To probe mechanisms of antagonism, we obtained receptor structures with the active component of curare, a poison arrow toxin and precursor to modern muscle relaxants. d-Tubocurarine stabilizes the receptor in a desensitized-like state in the presence and absence of agonist. These findings define the transitions between resting and desensitized states and reveal divergent means by which antagonists block channel activity of the muscle-type nicotinic receptor. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25206.map.gz emd_25206.map.gz | 5.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25206-v30.xml emd-25206-v30.xml emd-25206.xml emd-25206.xml | 18.9 KB 18.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25206_fsc.xml emd_25206_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_25206.png emd_25206.png | 90.1 KB | ||
| Others |  emd_25206_additional_1.map.gz emd_25206_additional_1.map.gz | 4.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25206 http://ftp.pdbj.org/pub/emdb/structures/EMD-25206 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25206 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25206 | HTTPS FTP |
-Validation report
| Summary document |  emd_25206_validation.pdf.gz emd_25206_validation.pdf.gz | 364.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25206_full_validation.pdf.gz emd_25206_full_validation.pdf.gz | 364.5 KB | Display | |
| Data in XML |  emd_25206_validation.xml.gz emd_25206_validation.xml.gz | 10.9 KB | Display | |
| Data in CIF |  emd_25206_validation.cif.gz emd_25206_validation.cif.gz | 14.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25206 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25206 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25206 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25206 | HTTPS FTP |
-Related structure data
| Related structure data |  7smrMC  7smmC  7smqC  7smsC  7smtC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25206.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25206.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of Torpedo acetylcholine receptor in complex with carbachol, desensitized state | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
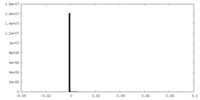
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Unsharpened map
| File | emd_25206_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
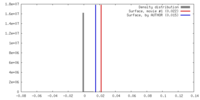
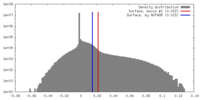
| Density Histograms |
- Sample components
Sample components
+Entire : Torpedo acetylcholine receptor in complex with carbachol, desensi...
+Supramolecule #1: Torpedo acetylcholine receptor in complex with carbachol, desensi...
+Macromolecule #1: Acetylcholine receptor subunit alpha
+Macromolecule #2: Acetylcholine receptor subunit delta
+Macromolecule #3: Acetylcholine receptor subunit beta
+Macromolecule #4: Acetylcholine receptor subunit gamma
+Macromolecule #8: 2-[(AMINOCARBONYL)OXY]-N,N,N-TRIMETHYLETHANAMINIUM
+Macromolecule #9: CHOLESTEROL
+Macromolecule #10: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
+Macromolecule #11: 2-acetamido-2-deoxy-beta-D-glucopyranose
+Macromolecule #12: nonane
+Macromolecule #13: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 35.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)