[English] 日本語
 Yorodumi
Yorodumi- EMDB-23526: Human TRiC/CCT complex with reovirus outer capsid protein sigma-3 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23526 | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human TRiC/CCT complex with reovirus outer capsid protein sigma-3 | |||||||||||||||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||||||||
 Keywords Keywords | tric / cct / reovirus capsid / sigma 3 / CHAPERONE | |||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationscaRNA localization to Cajal body / zona pellucida receptor complex / positive regulation of establishment of protein localization to telomere / symbiont-mediated suppression of host PKR/eIFalpha signaling / positive regulation of protein localization to Cajal body / chaperone mediated protein folding independent of cofactor / viral outer capsid / tubulin complex assembly / BBSome-mediated cargo-targeting to cilium / binding of sperm to zona pellucida ...scaRNA localization to Cajal body / zona pellucida receptor complex / positive regulation of establishment of protein localization to telomere / symbiont-mediated suppression of host PKR/eIFalpha signaling / positive regulation of protein localization to Cajal body / chaperone mediated protein folding independent of cofactor / viral outer capsid / tubulin complex assembly / BBSome-mediated cargo-targeting to cilium / binding of sperm to zona pellucida / positive regulation of telomerase RNA localization to Cajal body / Folding of actin by CCT/TriC / Formation of tubulin folding intermediates by CCT/TriC / chaperonin-containing T-complex / Prefoldin mediated transfer of substrate to CCT/TriC / protein serine/threonine kinase inhibitor activity / RHOBTB1 GTPase cycle / intermediate filament cytoskeleton / WD40-repeat domain binding / pericentriolar material / beta-tubulin binding / Association of TriC/CCT with target proteins during biosynthesis / : / chaperone-mediated protein complex assembly / RHOBTB2 GTPase cycle / heterochromatin / chaperone-mediated protein folding / positive regulation of telomere maintenance via telomerase / protein folding chaperone / viral life cycle / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / acrosomal vesicle / cell projection / mRNA 3'-UTR binding / ATP-dependent protein folding chaperone / response to virus / cilium / mRNA 5'-UTR binding / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / azurophil granule lumen / G-protein beta-subunit binding / unfolded protein binding / melanosome / protein folding / regulation of translation / cell body / secretory granule lumen / microtubule / ficolin-1-rich granule lumen / host cell cytoplasm / cytoskeleton / protein stabilization / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / symbiont-mediated suppression of host innate immune response / cadherin binding / virus-mediated perturbation of host defense response / centrosome / ubiquitin protein ligase binding / Neutrophil degranulation / host cell nucleus / structural molecule activity / Golgi apparatus / ATP hydrolysis activity / RNA binding / extracellular exosome / extracellular region / nucleoplasm / ATP binding / metal ion binding / cytosol / cytoplasm Similarity search - Function | |||||||||||||||||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Mammalian orthoreovirus 3 Dearing / Mammalian orthoreovirus 3 Dearing /  Reovirus type 3 (strain Dearing) Reovirus type 3 (strain Dearing) | |||||||||||||||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.2 Å | |||||||||||||||||||||||||||||||||
 Authors Authors | Knowlton JJ / Gestaut D / Ma B / Taylor G / Seven AB / Leitner A / Wilson GJ / Shanker S / Yates NA / Prasad BVV ...Knowlton JJ / Gestaut D / Ma B / Taylor G / Seven AB / Leitner A / Wilson GJ / Shanker S / Yates NA / Prasad BVV / Aebersold R / Chiu W / Frydman J / Dermody TS | |||||||||||||||||||||||||||||||||
| Funding support |  United States, European Union, 10 items United States, European Union, 10 items
| |||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2021 Journal: Proc Natl Acad Sci U S A / Year: 2021Title: Structural and functional dissection of reovirus capsid folding and assembly by the prefoldin-TRiC/CCT chaperone network. Authors: Jonathan J Knowlton / Daniel Gestaut / Boxue Ma / Gwen Taylor / Alpay Burak Seven / Alexander Leitner / Gregory J Wilson / Sreejesh Shanker / Nathan A Yates / B V Venkataram Prasad / Ruedi ...Authors: Jonathan J Knowlton / Daniel Gestaut / Boxue Ma / Gwen Taylor / Alpay Burak Seven / Alexander Leitner / Gregory J Wilson / Sreejesh Shanker / Nathan A Yates / B V Venkataram Prasad / Ruedi Aebersold / Wah Chiu / Judith Frydman / Terence S Dermody /   Abstract: Intracellular protein homeostasis is maintained by a network of chaperones that function to fold proteins into their native conformation. The eukaryotic TRiC chaperonin (TCP1-ring complex, also ...Intracellular protein homeostasis is maintained by a network of chaperones that function to fold proteins into their native conformation. The eukaryotic TRiC chaperonin (TCP1-ring complex, also called CCT for cytosolic chaperonin containing TCP1) facilitates folding of a subset of proteins with folding constraints such as complex topologies. To better understand the mechanism of TRiC folding, we investigated the biogenesis of an obligate TRiC substrate, the reovirus σ3 capsid protein. We discovered that the σ3 protein interacts with a network of chaperones, including TRiC and prefoldin. Using a combination of cryoelectron microscopy, cross-linking mass spectrometry, and biochemical approaches, we establish functions for TRiC and prefoldin in folding σ3 and promoting its assembly into higher-order oligomers. These studies illuminate the molecular dynamics of σ3 folding and establish a biological function for TRiC in virus assembly. In addition, our findings provide structural and functional insight into the mechanism by which TRiC and prefoldin participate in the assembly of protein complexes. | |||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23526.map.gz emd_23526.map.gz | 77.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23526-v30.xml emd-23526-v30.xml emd-23526.xml emd-23526.xml | 28.3 KB 28.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_23526.png emd_23526.png | 233 KB | ||
| Filedesc metadata |  emd-23526.cif.gz emd-23526.cif.gz | 9.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23526 http://ftp.pdbj.org/pub/emdb/structures/EMD-23526 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23526 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23526 | HTTPS FTP |
-Validation report
| Summary document |  emd_23526_validation.pdf.gz emd_23526_validation.pdf.gz | 600.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_23526_full_validation.pdf.gz emd_23526_full_validation.pdf.gz | 600 KB | Display | |
| Data in XML |  emd_23526_validation.xml.gz emd_23526_validation.xml.gz | 6.1 KB | Display | |
| Data in CIF |  emd_23526_validation.cif.gz emd_23526_validation.cif.gz | 6.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23526 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23526 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23526 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23526 | HTTPS FTP |
-Related structure data
| Related structure data |  7lupMC  7lumC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23526.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23526.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
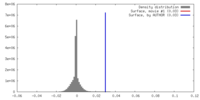
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : human TRiC/CCT complex with reovirus outer capsid protein sigma-3
+Supramolecule #1: human TRiC/CCT complex with reovirus outer capsid protein sigma-3
+Supramolecule #2: human TRiC/CCT complex
+Supramolecule #3: reovirus outer capsid protein sigma-3
+Macromolecule #1: T-complex protein 1 subunit epsilon
+Macromolecule #2: T-complex protein 1 subunit beta
+Macromolecule #3: T-complex protein 1 subunit delta
+Macromolecule #4: T-complex protein 1 subunit gamma
+Macromolecule #5: T-complex protein 1 subunit eta
+Macromolecule #6: T-complex protein 1 subunit theta
+Macromolecule #7: T-complex protein 1 subunit zeta
+Macromolecule #8: T-complex protein 1 subunit alpha
+Macromolecule #9: Outer capsid protein sigma-3
+Macromolecule #10: ZINC ION
+Macromolecule #11: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 36.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 6.2 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 26000 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)