[English] 日本語
 Yorodumi
Yorodumi- EMDB-23441: Structure of human SetD3 methyl-transferase in complex with 2A pr... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of human SetD3 methyl-transferase in complex with 2A protease from Coxsackievirus B3 | |||||||||
 Map data Map data | From cryosparc 2 non uniform refinement job, final map, filtered by FSC (3.5A), sharpened with b-factor of 180. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Methyltransferase / viral protease / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationprotein-histidine N-methyltransferase / peptidyl-histidine methylation / regulation of uterine smooth muscle contraction / protein-L-histidine N-tele-methyltransferase activity / actin modification / symbiont-mediated suppression of host NF-kappaB cascade / symbiont-mediated perturbation of host transcription / histone H3K36 methyltransferase activity / histone H3K4 methyltransferase activity / positive regulation of muscle cell differentiation ...protein-histidine N-methyltransferase / peptidyl-histidine methylation / regulation of uterine smooth muscle contraction / protein-L-histidine N-tele-methyltransferase activity / actin modification / symbiont-mediated suppression of host NF-kappaB cascade / symbiont-mediated perturbation of host transcription / histone H3K36 methyltransferase activity / histone H3K4 methyltransferase activity / positive regulation of muscle cell differentiation / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / PKMTs methylate histone lysines / endocytosis involved in viral entry into host cell / nucleoside-triphosphate phosphatase / protein complex oligomerization / monoatomic ion channel activity / actin binding / DNA replication / transcription coactivator activity / RNA polymerase II-specific DNA-binding transcription factor binding / RNA helicase activity / induction by virus of host autophagy / cysteine-type endopeptidase activity / RNA-directed RNA polymerase / viral RNA genome replication / virus-mediated perturbation of host defense response / RNA-dependent RNA polymerase activity / DNA-templated transcription / host cell nucleus / positive regulation of DNA-templated transcription / chromatin / virion attachment to host cell / structural molecule activity / positive regulation of transcription by RNA polymerase II / ATP hydrolysis activity / proteolysis / RNA binding / nucleoplasm / ATP binding / membrane / metal ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Coxsackievirus B3 (strain Nancy) / Coxsackievirus B3 (strain Nancy) /  Homo sapiens (human) Homo sapiens (human) | |||||||||
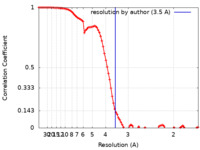
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Verba KA / Schulze-Gahmen U | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structure-function analysis of enterovirus protease 2A in complex with its essential host factor SETD3. Authors: Christine E Peters / Ursula Schulze-Gahmen / Manon Eckhardt / Gwendolyn M Jang / Jiewei Xu / Ernst H Pulido / Conner Bardine / Charles S Craik / Melanie Ott / Or Gozani / Kliment A Verba / ...Authors: Christine E Peters / Ursula Schulze-Gahmen / Manon Eckhardt / Gwendolyn M Jang / Jiewei Xu / Ernst H Pulido / Conner Bardine / Charles S Craik / Melanie Ott / Or Gozani / Kliment A Verba / Ruth Hüttenhain / Jan E Carette / Nevan J Krogan /  Abstract: Enteroviruses cause a number of medically relevant and widespread human diseases with no approved antiviral therapies currently available. Host-directed therapies present an enticing option for this ...Enteroviruses cause a number of medically relevant and widespread human diseases with no approved antiviral therapies currently available. Host-directed therapies present an enticing option for this diverse genus of viruses. We have previously identified the actin histidine methyltransferase SETD3 as a critical host factor physically interacting with the viral protease 2A. Here, we report the 3.5 Å cryo-EM structure of SETD3 interacting with coxsackievirus B3 2A at two distinct interfaces, including the substrate-binding surface within the SET domain. Structure-function analysis revealed that mutations of key residues in the SET domain resulted in severely reduced binding to 2A and complete protection from enteroviral infection. Our findings provide insight into the molecular basis of the SETD3-2A interaction and a framework for the rational design of host-directed therapeutics against enteroviruses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23441.map.gz emd_23441.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23441-v30.xml emd-23441-v30.xml emd-23441.xml emd-23441.xml | 20.8 KB 20.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_23441_fsc.xml emd_23441_fsc.xml | 9 KB | Display |  FSC data file FSC data file |
| Images |  emd_23441.png emd_23441.png | 137 KB | ||
| Masks |  emd_23441_msk_1.map emd_23441_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-23441.cif.gz emd-23441.cif.gz | 6.9 KB | ||
| Others |  emd_23441_half_map_1.map.gz emd_23441_half_map_1.map.gz emd_23441_half_map_2.map.gz emd_23441_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23441 http://ftp.pdbj.org/pub/emdb/structures/EMD-23441 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23441 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23441 | HTTPS FTP |
-Related structure data
| Related structure data |  7lmsMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23441.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23441.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | From cryosparc 2 non uniform refinement job, final map, filtered by FSC (3.5A), sharpened with b-factor of 180. | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.835 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_23441_msk_1.map emd_23441_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_23441_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_23441_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of Coxsackievirus B3 2A protease and human SetD3 ...
| Entire | Name: Ternary complex of Coxsackievirus B3 2A protease and human SetD3 methyltransferase |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of Coxsackievirus B3 2A protease and human SetD3 ...
| Supramolecule | Name: Ternary complex of Coxsackievirus B3 2A protease and human SetD3 methyltransferase type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Coxsackievirus B3 (strain Nancy) Coxsackievirus B3 (strain Nancy) |
| Molecular weight | Theoretical: 84 KDa |
-Macromolecule #1: Protease 2A
| Macromolecule | Name: Protease 2A / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: picornain 2A |
|---|---|
| Source (natural) | Organism:  Coxsackievirus B3 (strain Nancy) / Strain: Nancy Coxsackievirus B3 (strain Nancy) / Strain: Nancy |
| Molecular weight | Theoretical: 16.584482 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SNAMGQQSGA VYVGNYRVVN RHLATSADWQ NCVWESYNRD LLVSTTTAHG CDIIARCQCT TGVYFCASKN KHYPISFEGP GLVEVQESE YYPRRYQSHV LLAAGFSEPG DAGGILRCEH GVIGIVTMGG EGVVGFADIR DLLWLEDDAM EQ UniProtKB: Genome polyprotein |
-Macromolecule #2: Actin-histidine N-methyltransferase
| Macromolecule | Name: Actin-histidine N-methyltransferase / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: protein-histidine N-methyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 67.483117 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SNAGKKSRVK TQKSGTGATA TVSPKEILNL TSELLQKCSS PAPGPGKEWE EYVQIRTLVE KIRKKQKGLS VTFDGKREDY FPDLMKWAS ENGASVEGFE MVNFKEEGFG LRATRDIKAE ELFLWVPRKL LMTVESAKNS VLGPLYSQDR ILQAMGNIAL A FHLLCERA ...String: SNAGKKSRVK TQKSGTGATA TVSPKEILNL TSELLQKCSS PAPGPGKEWE EYVQIRTLVE KIRKKQKGLS VTFDGKREDY FPDLMKWAS ENGASVEGFE MVNFKEEGFG LRATRDIKAE ELFLWVPRKL LMTVESAKNS VLGPLYSQDR ILQAMGNIAL A FHLLCERA SPNSFWQPYI QTLPSEYDTP LYFEEDEVRY LQSTQAIHDV FSQYKNTARQ YAYFYKVIQT HPHANKLPLK DS FTYEDYR WAVSSVMTRQ NQIPTEDGSR VTLALIPLWD MCNHTNGLIT TGYNLEDDRC ECVALQDFRA GEQIYIFYGT RSN AEFVIH SGFFFDNNSH DRVKIKLGVS KSDRLYAMKA EVLARAGIPT SSVFALHFTE PPISAQLLAF LRVFCMTEEE LKEH LLGDS AIDRIFTLGN SEFPVSWDNE VKLWTFLEDR ASLLLKTYKT TIEEDKSVLK NHDLSVRAKM AIKLRLGEKE ILEKA VKSA AVNREYYRQQ MEEKAPLPKY EESNLGLLES SVGDSRLPLV LRNLEEEAGV QDALNIREAI SKAKATENGL VNGENS IPN GTRSENESLN QESKRAVEDA KGSSSDSTAG VKE UniProtKB: Actin-histidine N-methyltransferase |
-Macromolecule #3: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #4: S-ADENOSYL-L-HOMOCYSTEINE
| Macromolecule | Name: S-ADENOSYL-L-HOMOCYSTEINE / type: ligand / ID: 4 / Number of copies: 1 / Formula: SAH |
|---|---|
| Molecular weight | Theoretical: 384.411 Da |
| Chemical component information |  ChemComp-SAH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.34 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.2 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Details: 17mA | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 1200 / Average exposure time: 6.0 sec. / Average electron dose: 68.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-7lms: |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X