+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22808 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Myosin XI-F-actin complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | myosin / actin / cytoskeleton / motor protein / CONTRACTILE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationvesicle transport along actin filament / Striated Muscle Contraction / myosin complex / microfilament motor activity / skeletal muscle thin filament assembly / striated muscle thin filament / skeletal muscle fiber development / stress fiber / actin filament organization / actin filament ...vesicle transport along actin filament / Striated Muscle Contraction / myosin complex / microfilament motor activity / skeletal muscle thin filament assembly / striated muscle thin filament / skeletal muscle fiber development / stress fiber / actin filament organization / actin filament / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / actin filament binding / vesicle / calmodulin binding / hydrolase activity / ATP binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Chara corallina (plant) Chara corallina (plant) | |||||||||
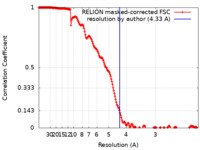
| Method | helical reconstruction / cryo EM / Resolution: 4.33 Å | |||||||||
 Authors Authors | Gong R / Alushin GM | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Chem Biol / Year: 2021 Journal: Nat Chem Biol / Year: 2021Title: Optical control of fast and processive engineered myosins in vitro and in living cells. Authors: Paul V Ruijgrok / Rajarshi P Ghosh / Sasha Zemsky / Muneaki Nakamura / Rui Gong / Lin Ning / Robert Chen / Vipul T Vachharajani / Alexander E Chu / Namrata Anand / Raphael R Eguchi / Po-Ssu ...Authors: Paul V Ruijgrok / Rajarshi P Ghosh / Sasha Zemsky / Muneaki Nakamura / Rui Gong / Lin Ning / Robert Chen / Vipul T Vachharajani / Alexander E Chu / Namrata Anand / Raphael R Eguchi / Po-Ssu Huang / Michael Z Lin / Gregory M Alushin / Jan T Liphardt / Zev Bryant /  Abstract: Precision tools for spatiotemporal control of cytoskeletal motor function are needed to dissect fundamental biological processes ranging from intracellular transport to cell migration and division. ...Precision tools for spatiotemporal control of cytoskeletal motor function are needed to dissect fundamental biological processes ranging from intracellular transport to cell migration and division. Direct optical control of motor speed and direction is one promising approach, but it remains a challenge to engineer controllable motors with desirable properties such as the speed and processivity required for transport applications in living cells. Here, we develop engineered myosin motors that combine large optical modulation depths with high velocities, and create processive myosin motors with optically controllable directionality. We characterize the performance of the motors using in vitro motility assays, single-molecule tracking and live-cell imaging. Bidirectional processive motors move efficiently toward the tips of cellular protrusions in the presence of blue light, and can transport molecular cargo in cells. Robust gearshifting myosins will further enable programmable transport in contexts ranging from in vitro active matter reconstitutions to microfabricated systems that harness molecular propulsion. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22808.map.gz emd_22808.map.gz | 282.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22808-v30.xml emd-22808-v30.xml emd-22808.xml emd-22808.xml | 20.8 KB 20.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22808_fsc.xml emd_22808_fsc.xml | 18.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_22808.png emd_22808.png | 72.4 KB | ||
| Masks |  emd_22808_msk_1.map emd_22808_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-22808.cif.gz emd-22808.cif.gz | 6.9 KB | ||
| Others |  emd_22808_half_map_1.map.gz emd_22808_half_map_1.map.gz emd_22808_half_map_2.map.gz emd_22808_half_map_2.map.gz | 411.1 MB 411 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22808 http://ftp.pdbj.org/pub/emdb/structures/EMD-22808 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22808 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22808 | HTTPS FTP |
-Validation report
| Summary document |  emd_22808_validation.pdf.gz emd_22808_validation.pdf.gz | 933.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22808_full_validation.pdf.gz emd_22808_full_validation.pdf.gz | 932.8 KB | Display | |
| Data in XML |  emd_22808_validation.xml.gz emd_22808_validation.xml.gz | 26.6 KB | Display | |
| Data in CIF |  emd_22808_validation.cif.gz emd_22808_validation.cif.gz | 35.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22808 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22808 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22808 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22808 | HTTPS FTP |
-Related structure data
| Related structure data |  7kchMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22808.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22808.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.096 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_22808_msk_1.map emd_22808_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_22808_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_22808_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Myosin XI-F-actin complex
| Entire | Name: Myosin XI-F-actin complex |
|---|---|
| Components |
|
-Supramolecule #1: Myosin XI-F-actin complex
| Supramolecule | Name: Myosin XI-F-actin complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 209 kDa/nm |
-Macromolecule #1: Actin, alpha skeletal muscle
| Macromolecule | Name: Actin, alpha skeletal muscle / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 42.096953 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MCDEDETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIEHGIITN WDDMEKIWH HTFYNELRVA PEEHPTLLTE APLNPKANRE KMTQIMFETF NVPAMYVAIQ AVLSLYASGR TTGIVLDSGD G VTHNVPIY ...String: MCDEDETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIEHGIITN WDDMEKIWH HTFYNELRVA PEEHPTLLTE APLNPKANRE KMTQIMFETF NVPAMYVAIQ AVLSLYASGR TTGIVLDSGD G VTHNVPIY EGYALPHAIM RLDLAGRDLT DYLMKILTER GYSFVTTAER EIVRDIKEKL CYVALDFENE MATAASSSSL EK SYELPDG QVITIGNERF RCPETLFQPS FIGMESAGIH ETTYNSIMKC DIDIRKDLYA NNVMSGGTTM YPGIADRMQK EIT ALAPST MKIKIIAPPE RKYSVWIGGS ILASLSTFQQ MWITKQEYDE AGPSIVHRKC F UniProtKB: Actin, alpha skeletal muscle |
-Macromolecule #2: Unconventional myosin heavy chain
| Macromolecule | Name: Unconventional myosin heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chara corallina (plant) Chara corallina (plant) |
| Molecular weight | Theoretical: 83.01532 KDa |
| Recombinant expression | Organism:  Spodoptera aff. frugiperda 2 RZ-2014 (butterflies/moths) Spodoptera aff. frugiperda 2 RZ-2014 (butterflies/moths) |
| Sequence | String: GSPAWVEDVE TVWIEATVVK LDGDAITART VNGDLVETTM ANALPRDEDV TMRGVDDMTK LSYLHEPGVL HNLYTRFKHD EIYTFTGNI LIAVNPFTRL PHLFNTYMMK QYQDAQPGDL NPHVYSVADA AYKAMMEEMK SQAILVSGES GAGKTETTKQ I MQYLAFVG ...String: GSPAWVEDVE TVWIEATVVK LDGDAITART VNGDLVETTM ANALPRDEDV TMRGVDDMTK LSYLHEPGVL HNLYTRFKHD EIYTFTGNI LIAVNPFTRL PHLFNTYMMK QYQDAQPGDL NPHVYSVADA AYKAMMEEMK SQAILVSGES GAGKTETTKQ I MQYLAFVG GRTVGDERSV EQQVLQSNPL LEAFGNAKTV RNNNSSRFGK FVEIQFNNGK ISGAAVRTYL LERSRVTQIS SP ERNYHCF YQLVAGASPE DAERLKLGPP DSFHYLNQSK CVEVGAIDDC KEYQLTREAM DIVGITTEEQ EAIFRTIAAV LHL GNIEFD SGESDASEVS TEKSKFHLKA AAEMLMCDEQ MLEKSLTTRI MKATRTESIT KILNKSQATD NRDSIAKTIY AKLF DWLVN KVNKSIGQDP HSTVLIGVLD IYGFESFEIN SFEQFCINLT NEKLQQHFNT HVFKMEQAEY RKEEINWDNI DFVDN IDVL DLIEKKPLGI IALLDEACML PRSTAESFAR KLGDTFNNHR RFSKHKFKRT AFTIDHYAGQ VEYRADLFLE KNKDFV VPE HQQLLHASRC AFVSGLFPAD EGGGKKGGTK APSKKKFMSI GSQFKLQLAA LMETLKLTAP HYIRCVKPNM QLKPQIF EN KNVLQQLRCS GVLEAVRISC AGFPTRRTFE EFLDRFGLLH PEVLIESAEE SADEKVACQN LLEKCNLKGY QIGKTKVF L RAGQMAILDT LRSN UniProtKB: Unconventional myosin heavy chain |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 3 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 3 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7 Component:
Details: 10 mM imidazole pH 7.0,50 mM KCl,1mM MgCl2, 1mM EGTA, 0.5 mM DTT, 0.01% NaN3 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 20 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 10 sec. / Pretreatment - Atmosphere: OTHER | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 67.12 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.0 mm / Nominal defocus max: -4.0 µm / Nominal defocus min: -1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)