[English] 日本語
 Yorodumi
Yorodumi- EMDB-20349: VC-Tn6677 multisubunit CRISPR/Cas effector, close conformation. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20349 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | VC-Tn6677 multisubunit CRISPR/Cas effector, close conformation. | |||||||||
 Map data Map data | Post-processed map with automatically determined B-factor by Relion. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CRISPR/Cas / Cascade / RNA BINDING PROTEIN / RNA BINDING PROTEIN-RNA complex | |||||||||
| Biological species |  | |||||||||
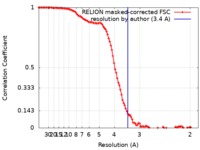
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Halpin-Healy T / Klompe S | |||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Structural basis of DNA targeting by a transposon-encoded CRISPR-Cas system. Authors: Tyler S Halpin-Healy / Sanne E Klompe / Samuel H Sternberg / Israel S Fernández /  Abstract: Bacteria use adaptive immune systems encoded by CRISPR and Cas genes to maintain genomic integrity when challenged by pathogens and mobile genetic elements. Type I CRISPR-Cas systems typically ...Bacteria use adaptive immune systems encoded by CRISPR and Cas genes to maintain genomic integrity when challenged by pathogens and mobile genetic elements. Type I CRISPR-Cas systems typically target foreign DNA for degradation via joint action of the ribonucleoprotein complex Cascade and the helicase-nuclease Cas3, but nuclease-deficient type I systems lacking Cas3 have been repurposed for RNA-guided transposition by bacterial Tn7-like transposons. How CRISPR- and transposon-associated machineries collaborate during DNA targeting and insertion remains unknown. Here we describe structures of a TniQ-Cascade complex encoded by the Vibrio cholerae Tn6677 transposon using cryo-electron microscopy, revealing the mechanistic basis of this functional coupling. The cryo-electron microscopy maps enabled de novo modelling and refinement of the transposition protein TniQ, which binds to the Cascade complex as a dimer in a head-to-tail configuration, at the interface formed by Cas6 and Cas7 near the 3' end of the CRISPR RNA (crRNA). The natural Cas8-Cas5 fusion protein binds the 5' crRNA handle and contacts the TniQ dimer via a flexible insertion domain. A target DNA-bound structure reveals critical interactions necessary for protospacer-adjacent motif recognition and R-loop formation. This work lays the foundation for a structural understanding of how DNA targeting by TniQ-Cascade leads to downstream recruitment of additional transposase proteins, and will guide protein engineering efforts to leverage this system for programmable DNA insertions in genome-engineering applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20349.map.gz emd_20349.map.gz | 96.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20349-v30.xml emd-20349-v30.xml emd-20349.xml emd-20349.xml | 29.3 KB 29.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20349_fsc.xml emd_20349_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_20349.png emd_20349.png | 184.9 KB | ||
| Masks |  emd_20349_msk_1.map emd_20349_msk_1.map emd_20349_msk_2.map emd_20349_msk_2.map emd_20349_msk_3.map emd_20349_msk_3.map | 103 MB 103 MB 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-20349.cif.gz emd-20349.cif.gz | 6.9 KB | ||
| Others |  emd_20349_additional_1.map.gz emd_20349_additional_1.map.gz emd_20349_additional_2.map.gz emd_20349_additional_2.map.gz emd_20349_additional_3.map.gz emd_20349_additional_3.map.gz emd_20349_additional_4.map.gz emd_20349_additional_4.map.gz emd_20349_half_map_1.map.gz emd_20349_half_map_1.map.gz emd_20349_half_map_2.map.gz emd_20349_half_map_2.map.gz | 11.4 MB 8.1 MB 8.3 MB 80.7 MB 81.1 MB 81.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20349 http://ftp.pdbj.org/pub/emdb/structures/EMD-20349 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20349 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20349 | HTTPS FTP |
-Related structure data
| Related structure data |  6pifMC  6pigC  6pijC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20349.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20349.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed map with automatically determined B-factor by Relion. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.95 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_20349_msk_1.map emd_20349_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Mask #2
| File |  emd_20349_msk_2.map emd_20349_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Mask #3
| File |  emd_20349_msk_3.map emd_20349_msk_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Post-processed map with automatically determined B-factor by Relion...
| File | emd_20349_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed map with automatically determined B-factor by Relion masked with the the post-processing mask. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Post-processed map focused on cas5 8, cas7.6-4
| File | emd_20349_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed map focused on cas5_8, cas7.6-4 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Post-processed map focused on TniQ dimer, cas6. cas7.1...
| File | emd_20349_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed map focused on TniQ dimer, cas6. cas7.1 and the RNA handle | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened map
| File | emd_20349_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Unsharpened half map 1 from Relion3 AutoRefine
| File | emd_20349_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened half map 1 from Relion3 AutoRefine | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Unsharpened half map 2 from Relion3 AutoRefine
| File | emd_20349_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened half map 2 from Relion3 AutoRefine | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : VC-Tn667 transposon encoded CRISPR/Cas effector
| Entire | Name: VC-Tn667 transposon encoded CRISPR/Cas effector |
|---|---|
| Components |
|
-Supramolecule #1: VC-Tn667 transposon encoded CRISPR/Cas effector
| Supramolecule | Name: VC-Tn667 transposon encoded CRISPR/Cas effector / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 600 KDa |
-Macromolecule #1: Cas7, type I-F CRISPR-associated protein
| Macromolecule | Name: Cas7, type I-F CRISPR-associated protein / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 39.517586 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KLPTNLAYER SIDPSDVCFF VVWPDDRKTP LTYNSRTLLG QMEAKSLAYD VSGQPIKSAT AEALAQGNPH QVDFCHVPYG ASHIECSFS VSFSSELRQP YKCNSSKVKQ TLVQLVELYE TKIGWTELAT RYLMNICNGK WLWKNTRKAY CWNIVLTPWP W NGEKVGFE ...String: KLPTNLAYER SIDPSDVCFF VVWPDDRKTP LTYNSRTLLG QMEAKSLAYD VSGQPIKSAT AEALAQGNPH QVDFCHVPYG ASHIECSFS VSFSSELRQP YKCNSSKVKQ TLVQLVELYE TKIGWTELAT RYLMNICNGK WLWKNTRKAY CWNIVLTPWP W NGEKVGFE DIRTNYTSRQ DFKNNKNWSA IVEMIKTAFS STDGLAIFEV RATLHLPTNA MVRPSQVFTE KEAAAAAAAT QN SRVFQST TIDGERSPIL GAFKTGAAIA TIDDWYPEAT EPLRVGRFGV HREDVTCYRH PSTGKDFFSI LQQAEHYIEV LSA NKTPAQ ETINDMHFLM ANLIKGGMFQ HKGD |
-Macromolecule #2: cas5_8 naturally occurring fusion protein
| Macromolecule | Name: cas5_8 naturally occurring fusion protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 58.26698 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: LKELIASNPD DLTTELKRAF RPLTPHIAID GNELDALTIL VNLTDKTDDQ KDLLDRAKCK QKLRDEKWWA SCINCVNYRQ SHNPKFPDI RSEGVIRTQA LGELPSFLLS SSKIPPYHWS YSHDSKYVNK SAFLTNEFCW DGEISCLGEL LKDADHPLWN T LKKLGCSQ ...String: LKELIASNPD DLTTELKRAF RPLTPHIAID GNELDALTIL VNLTDKTDDQ KDLLDRAKCK QKLRDEKWWA SCINCVNYRQ SHNPKFPDI RSEGVIRTQA LGELPSFLLS SSKIPPYHWS YSHDSKYVNK SAFLTNEFCW DGEISCLGEL LKDADHPLWN T LKKLGCSQ KTCKAMAKQL ADITLTTINV TLAPNYLTQI SLPDSDTSYI SLSPVASLSM QSHFHQRLQD ENRHSAITRF SR TTNMGVT AMTCGGAFRM LKSGAKFSSP PHHRLNNGSF LVLPNIRVCG ATALSSPVTV GIPSLTAFFG FVHAFERNIN RTT SSFRVE SFAICVHQLH VEKRGLTAEF VEKGDGTISA PATRDDWQCD VVFSLILNTN FAQHIDQDTL VTSLPKRLAR GSAK IAIDD FKHINSFSTL ETAIESLPIE AGRWLSLYAQ SNNNLSDLLA AMTEDHQLMA SCVGYHLLEE PKDKPNSLRG YKHAI AECI IGLINSITFS SETDPNTIFW SLKNYQNYLV VQPRSIN |
-Macromolecule #3: type I-F CRISPR-associated endoribonuclease Cas6/Csy4
| Macromolecule | Name: type I-F CRISPR-associated endoribonuclease Cas6/Csy4 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.999234 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KWYYKTITFL PELCNNESLA AKCLRVLHGF NYQYETRNIG VSFPLWCDAT VGKKISFVSK NKIELDLLLK QHYFVQMEQL QYFHISNTV LVPEDCTYVS FRRCQSIDKL TAAGLARKIR RLEKRALSRG EQFDPSSFAQ KEHTAIAHYH SLGESSKQTN R NFRLNIRM ...String: KWYYKTITFL PELCNNESLA AKCLRVLHGF NYQYETRNIG VSFPLWCDAT VGKKISFVSK NKIELDLLLK QHYFVQMEQL QYFHISNTV LVPEDCTYVS FRRCQSIDKL TAAGLARKIR RLEKRALSRG EQFDPSSFAQ KEHTAIAHYH SLGESSKQTN R NFRLNIRM LSEQPREGNS IFSSYGLSNS ENSFQPVPLI |
-Macromolecule #4: TniQ monomer 1
| Macromolecule | Name: TniQ monomer 1 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 41.51716 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AMFLQRPKPY SDESLESFFI RVANKNGYGD VHRFLEATKR FLQDIDHNGY QTFPTDITRI NPYSAKNSSS ARTASFLKLA QLTFNEPPE LLGLAINRTN MKYSPSTSAV VRGAEVFPRS LLRTHSIPCC PLCLRENGYA SYLWHFQGYE YCHSHNVPLI T TCSGHEAA ...String: AMFLQRPKPY SDESLESFFI RVANKNGYGD VHRFLEATKR FLQDIDHNGY QTFPTDITRI NPYSAKNSSS ARTASFLKLA QLTFNEPPE LLGLAINRTN MKYSPSTSAV VRGAEVFPRS LLRTHSIPCC PLCLRENGYA SYLWHFQGYE YCHSHNVPLI T TCSGHEAA CTVSNWLAGH ESKPLPNLPK SYRWGLVHWW MGIKDSDHFS FVQFFSNWPR SFHSIIEDEV EFNLEHAVVS TS ELRLKDL LGRLFFGSIR LPERNLQHNI ILGELLCYLE NRLWQDKGLI ANLKMNALEA TVMLNCSLDQ IASMVEQRIL KPN RKSKDV TDYLFHFGDI FCLWLAEFQS DEFNRSFYVS RW |
-Macromolecule #5: TniQ monomer 2
| Macromolecule | Name: TniQ monomer 2 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 42.373066 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AMFLQRPKPY SDESLESFFI RVANKNGYGD VHRFLEATKR FLQDIDHNGY QTFPTDITRI NPYSAKNSSS ARTASFLKLA QLTFNEPPE LLGLAINRTN MKYSPSTSAV VRGAEVFPRS LLRTHSIPCC PLCLRENGYA SYLWHFQGYE YCHSHNVPLI T TCSCGKEF ...String: AMFLQRPKPY SDESLESFFI RVANKNGYGD VHRFLEATKR FLQDIDHNGY QTFPTDITRI NPYSAKNSSS ARTASFLKLA QLTFNEPPE LLGLAINRTN MKYSPSTSAV VRGAEVFPRS LLRTHSIPCC PLCLRENGYA SYLWHFQGYE YCHSHNVPLI T TCSCGKEF DYRVSEAACT VSNWLAGHES KPLPNLPKSY RWGLVHWWMG IKDDHFSFVQ FFSNWPRSFH SIIEDEVEFN LE HAVVSTS ELRLKDLLGR LFFGSIRLPE RNLQHNIILG ELLCYLENRL WQDKGLIANL KMNALEATVM LNCSLDQIAS MVE QRILKP NAAAAAAAAA DVTDYLFHFG DIFCLWLAEF QSDEFNRSFY VSR |
-Macromolecule #6: guide RNA (60-MER)
| Macromolecule | Name: guide RNA (60-MER) / type: rna / ID: 6 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 19.237338 KDa |
| Sequence | String: CUGAUAACUU ACAGGACGCU UUGGCUUCAU UGCUUUUCAG GUGAACUGCC GAGUAGGUAG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: UltrAuFoil / Material: GOLD / Mesh: 300 | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 4 K / Instrument: FEI VITROBOT MARK II |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)