[English] 日本語
 Yorodumi
Yorodumi- EMDB-20212: Cryo-EM structure of Bimetallic dodecameric cage design 3 (BMC3) ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20212 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Bimetallic dodecameric cage design 3 (BMC3) from cytochrome cb562 | |||||||||||||||
 Map data Map data | 2.6 A map of dodecameric cage | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Supramolecular assembly / protein cage / bimetallic / metal binding / hydroxamic acid / METAL BINDING PROTEIN | |||||||||||||||
| Function / homology | Cytochrome b562 / Cytochrome b562 / Cytochrome c/b562 / electron transport chain / periplasmic space / electron transfer activity / iron ion binding / heme binding / Soluble cytochrome b562 Function and homology information Function and homology information | |||||||||||||||
| Biological species |  | |||||||||||||||
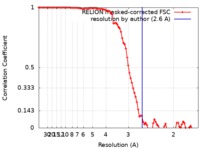
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||||||||
 Authors Authors | Golub E / Subramanian RH | |||||||||||||||
| Funding support |  United States, European Union, United States, European Union,  Germany, 4 items Germany, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Constructing protein polyhedra via orthogonal chemical interactions. Authors: Eyal Golub / Rohit H Subramanian / Julian Esselborn / Robert G Alberstein / Jake B Bailey / Jerika A Chiong / Xiaodong Yan / Timothy Booth / Timothy S Baker / F Akif Tezcan /  Abstract: Many proteins exist naturally as symmetrical homooligomers or homopolymers. The emergent structural and functional properties of such protein assemblies have inspired extensive efforts in ...Many proteins exist naturally as symmetrical homooligomers or homopolymers. The emergent structural and functional properties of such protein assemblies have inspired extensive efforts in biomolecular design. As synthesized by ribosomes, proteins are inherently asymmetric. Thus, they must acquire multiple surface patches that selectively associate to generate the different symmetry elements needed to form higher-order architectures-a daunting task for protein design. Here we address this problem using an inorganic chemical approach, whereby multiple modes of protein-protein interactions and symmetry are simultaneously achieved by selective, 'one-pot' coordination of soft and hard metal ions. We show that a monomeric protein (protomer) appropriately modified with biologically inspired hydroxamate groups and zinc-binding motifs assembles through concurrent Fe and Zn coordination into discrete dodecameric and hexameric cages. Our cages closely resemble natural polyhedral protein architectures and are, to our knowledge, unique among designed systems in that they possess tightly packed shells devoid of large apertures. At the same time, they can assemble and disassemble in response to diverse stimuli, owing to their heterobimetallic construction on minimal interprotein-bonding footprints. With stoichiometries ranging from [2 Fe:9 Zn:6 protomers] to [8 Fe:21 Zn:12 protomers], these protein cages represent some of the compositionally most complex protein assemblies-or inorganic coordination complexes-obtained by design. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20212.map.gz emd_20212.map.gz | 56.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20212-v30.xml emd-20212-v30.xml emd-20212.xml emd-20212.xml | 18.5 KB 18.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20212_fsc.xml emd_20212_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_20212.png emd_20212.png | 82.2 KB | ||
| Filedesc metadata |  emd-20212.cif.gz emd-20212.cif.gz | 6.4 KB | ||
| Others |  emd_20212_half_map_1.map.gz emd_20212_half_map_1.map.gz emd_20212_half_map_2.map.gz emd_20212_half_map_2.map.gz | 49.8 MB 49.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20212 http://ftp.pdbj.org/pub/emdb/structures/EMD-20212 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20212 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20212 | HTTPS FTP |
-Related structure data
| Related structure data |  6ovhMC  6ot4C  6ot7C  6ot8C  6ot9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20212.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20212.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 2.6 A map of dodecameric cage | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8452 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: half-map 2 prior to postprocessing in Relion 3.0
| File | emd_20212_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map 2 prior to postprocessing in Relion 3.0 | ||||||||||||
| Projections & Slices |
| ||||||||||||
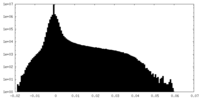
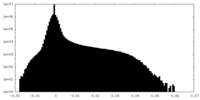
| Density Histograms |
-Half map: half-map 1 prior to postprocessing in Relion 3.0
| File | emd_20212_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map 1 prior to postprocessing in Relion 3.0 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Bimetallic dodecameric cage 3 (BMC3)
| Entire | Name: Bimetallic dodecameric cage 3 (BMC3) |
|---|---|
| Components |
|
-Supramolecule #1: Bimetallic dodecameric cage 3 (BMC3)
| Supramolecule | Name: Bimetallic dodecameric cage 3 (BMC3) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: Cryo-EM reconstruction of self-assembled BMC3 dodecameric cages |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 150 KDa |
-Macromolecule #1: Soluble cytochrome b562
| Macromolecule | Name: Soluble cytochrome b562 / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.809307 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ADLEHNMHTL NDNLKHIEKA DNATTVKDAL TKMQAAAQDA WSATPPKLED KSPDSPEMSD FRCGFWELIG QINAALHLAK QCKVKEAQA AAEQLKTTCN ACHQKYR UniProtKB: Soluble cytochrome b562 |
-Macromolecule #2: HEME C
| Macromolecule | Name: HEME C / type: ligand / ID: 2 / Number of copies: 12 / Formula: HEC |
|---|---|
| Molecular weight | Theoretical: 618.503 Da |
| Chemical component information |  ChemComp-HEC: |
-Macromolecule #3: ACETOHYDROXAMIC ACID
| Macromolecule | Name: ACETOHYDROXAMIC ACID / type: ligand / ID: 3 / Number of copies: 24 / Formula: HAE |
|---|---|
| Molecular weight | Theoretical: 75.067 Da |
| Chemical component information |  ChemComp-HAE: |
-Macromolecule #4: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 4 / Number of copies: 24 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #5: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 5 / Number of copies: 8 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 171 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.5 / Component - Concentration: 20.0 millimolar / Component - Name: Tris |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 101.325 kPa / Details: The grid was glow discharged at 20 mA for 30 s. |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
| Details | Protein solutions containing 20 micromolar BMC3 in 20 mM Tris (pH 8.5) were incubated with [FeSO4] = 20 micromolar, [ZnCl2] = 60 micromolar for 2-3 h to form cages. Samples were concentrated 10 fold prior to grid preparation. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number real images: 4672 / Average exposure time: 10.0 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | Symmetry mates were generated from the atomic model to build the dodecameric cage. Chain IDs were re-assigned to ascend from A-L. The cage PDB was manually fit to the EM density map in UCSF Chimera and refined using phenix.real_space_refine |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-6ovh: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)