[English] 日本語
 Yorodumi
Yorodumi- EMDB-17702: Cas9 bound to cognate DNA, Streptococcus thermophilus DGCC 7710 C... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cas9 bound to cognate DNA, Streptococcus thermophilus DGCC 7710 CRISPR3 system | |||||||||
 Map data Map data | Map sharpened using phenix.auto_sharpen_1.21rc1-4903 using _iso_to_d_cut procedure and 3.24A cutoff. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cas9 / CRISPR-Cas / PAM / spacer acquisition / RNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmaintenance of CRISPR repeat elements / endonuclease activity / defense response to virus / Hydrolases; Acting on ester bonds / DNA binding / RNA binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Streptococcus thermophilus DGCC 7710 (bacteria) / synthetic construct (others) Streptococcus thermophilus DGCC 7710 (bacteria) / synthetic construct (others) | |||||||||
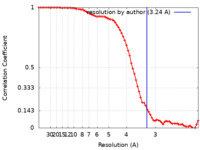
| Method | single particle reconstruction / cryo EM / Resolution: 3.24 Å | |||||||||
 Authors Authors | Sasnauskas G / Gaizauskaite U / Tamulaitiene G | |||||||||
| Funding support | Lithuania, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural basis for spacer acquisition in a type II-A CRISPR-Cas system Authors: Sasnauskas G / Gaizauskaite U / Tamulaitiene G | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17702.map.gz emd_17702.map.gz | 27.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17702-v30.xml emd-17702-v30.xml emd-17702.xml emd-17702.xml | 23.7 KB 23.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17702_fsc.xml emd_17702_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_17702.png emd_17702.png | 157.1 KB | ||
| Masks |  emd_17702_msk_1.map emd_17702_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17702.cif.gz emd-17702.cif.gz | 7.3 KB | ||
| Others |  emd_17702_additional_1.map.gz emd_17702_additional_1.map.gz emd_17702_half_map_1.map.gz emd_17702_half_map_1.map.gz emd_17702_half_map_2.map.gz emd_17702_half_map_2.map.gz | 15.3 MB 28.3 MB 28.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17702 http://ftp.pdbj.org/pub/emdb/structures/EMD-17702 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17702 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17702 | HTTPS FTP |
-Validation report
| Summary document |  emd_17702_validation.pdf.gz emd_17702_validation.pdf.gz | 882.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_17702_full_validation.pdf.gz emd_17702_full_validation.pdf.gz | 881.8 KB | Display | |
| Data in XML |  emd_17702_validation.xml.gz emd_17702_validation.xml.gz | 12.7 KB | Display | |
| Data in CIF |  emd_17702_validation.cif.gz emd_17702_validation.cif.gz | 17.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17702 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17702 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17702 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17702 | HTTPS FTP |
-Related structure data
| Related structure data |  8pj9MC  8pk1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17702.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17702.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map sharpened using phenix.auto_sharpen_1.21rc1-4903 using _iso_to_d_cut procedure and 3.24A cutoff. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17702_msk_1.map emd_17702_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
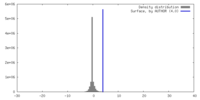
| Density Histograms |
-Additional map: unsharpened map
| File | emd_17702_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
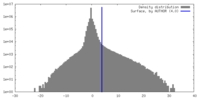
| Density Histograms |
-Half map: #1
| File | emd_17702_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
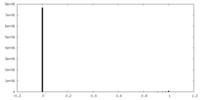
| Density Histograms |
-Half map: #2
| File | emd_17702_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
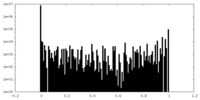
| Density Histograms |
- Sample components
Sample components
+Entire : Complex of Cas9 bound to crRNA:tracrRNA and to a cognate DNA duplex
+Supramolecule #1: Complex of Cas9 bound to crRNA:tracrRNA and to a cognate DNA duplex
+Supramolecule #2: CRISPR-associated endonuclease Cas9
+Supramolecule #3: crRNA, chain B and tracrRNA, chain C
+Supramolecule #4: DNA oligoduplex, target strand, chains D,E and DNA oligoduplex, n...
+Macromolecule #1: CRISPR-associated endonuclease Cas9
+Macromolecule #2: crRNA, chain B
+Macromolecule #3: tracrRNA, chain C
+Macromolecule #4: DNA oligoduplex, target strand, chains D,E
+Macromolecule #5: DNA oligoduplex, non-target strand, chain F
+Macromolecule #6: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 1744 / Average exposure time: 46.33 sec. / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 92000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Output model |  PDB-8pj9: |
 Movie
Movie Controller
Controller





 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)