+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Caenorhabditis elegans DPF-3 (apo) | |||||||||||||||
 Map data Map data | Full map processed with LocScale. | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Dipeptidylpeptidase / HYDROLASE | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationdipeptidyl-peptidase activity / serine-type peptidase activity / proteolysis Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
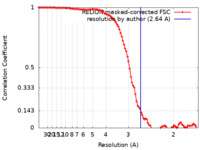
| Method | single particle reconstruction / cryo EM / Resolution: 2.64 Å | |||||||||||||||
 Authors Authors | Gudipati RK / Cavadini S / Kempf G / Grosshans H | |||||||||||||||
| Funding support |  Switzerland, Switzerland,  Poland, 4 items Poland, 4 items
| |||||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of Caenorhabditis elegans DPF-3 (apo) Authors: Gudipati RK / Cavadini S / Kempf G / Grosshans H | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17582.map.gz emd_17582.map.gz | 41.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17582-v30.xml emd-17582-v30.xml emd-17582.xml emd-17582.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17582_fsc.xml emd_17582_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_17582.png emd_17582.png | 119.2 KB | ||
| Filedesc metadata |  emd-17582.cif.gz emd-17582.cif.gz | 5.8 KB | ||
| Others |  emd_17582_additional_1.map.gz emd_17582_additional_1.map.gz emd_17582_half_map_1.map.gz emd_17582_half_map_1.map.gz emd_17582_half_map_2.map.gz emd_17582_half_map_2.map.gz | 64 MB 64.1 MB 64.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17582 http://ftp.pdbj.org/pub/emdb/structures/EMD-17582 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17582 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17582 | HTTPS FTP |
-Validation report
| Summary document |  emd_17582_validation.pdf.gz emd_17582_validation.pdf.gz | 751.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_17582_full_validation.pdf.gz emd_17582_full_validation.pdf.gz | 751.1 KB | Display | |
| Data in XML |  emd_17582_validation.xml.gz emd_17582_validation.xml.gz | 17.2 KB | Display | |
| Data in CIF |  emd_17582_validation.cif.gz emd_17582_validation.cif.gz | 22.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17582 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17582 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17582 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17582 | HTTPS FTP |
-Related structure data
| Related structure data |  8pbaMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17582.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17582.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map processed with LocScale. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.845 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Full map.
| File | emd_17582_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
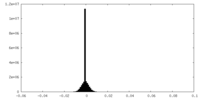
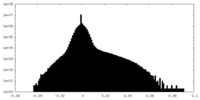
| Density Histograms |
-Half map: Half map 2.
| File | emd_17582_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1.
| File | emd_17582_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Homodimeric complex of DPF-3
| Entire | Name: Homodimeric complex of DPF-3 |
|---|---|
| Components |
|
-Supramolecule #1: Homodimeric complex of DPF-3
| Supramolecule | Name: Homodimeric complex of DPF-3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Dipeptidyl Peptidase Four (IV) family
| Macromolecule | Name: Dipeptidyl Peptidase Four (IV) family / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 106.462812 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MMFNFYQFLY NLQNVSPFID FSVLKQLTHT KMRENEPARF ETRSFSQLID HARSWKTEVR GMTTQGFTKI SLMRAEKDRL NMYAISSVP GTNTQSIFSV TIPLELVEKA QVADRKFELK LKSGYNVDSY IRKTPPSAEF TLQCERQRSQ VVTGISDYEI R NGKMILMA ...String: MMFNFYQFLY NLQNVSPFID FSVLKQLTHT KMRENEPARF ETRSFSQLID HARSWKTEVR GMTTQGFTKI SLMRAEKDRL NMYAISSVP GTNTQSIFSV TIPLELVEKA QVADRKFELK LKSGYNVDSY IRKTPPSAEF TLQCERQRSQ VVTGISDYEI R NGKMILMA GDQLFRYNPL NEALAAIPIA VPDDQSSTEP MDISEGSITS GTKGSGSEAP QSSTVPPVTR IPIKKPTTST EK PATAPPT NNFVSSAKVC PADSSLLAYV LNKQVYIEKN GKIIHRTSSN SKHITNGVPS YIVQEELERF EGIWWSESKT RLL YEHVNE EKVAESQFGV NGDPPVAPMK YPRAGTKNAY STLRMVILEN GKAYDVPLKD EVIYKHCPFY EYITRAGFFS DGTT VWVQV MSRDQAQCSL LLIPYTDFLL PEELGGSIKE DNLQLSTDLN MGVWDDKSHE ETMEKPPRGK LRGTVQIHKA RNDYW INTH NAIYPLKITD EEHPMYEFIY CLEKPNGSCL ALISAELDQN GYCRHTEEKL LMAENFSINK SMGIVVDEVR ELVYYV ANE SHPTEWNICV SHYRTGQHAQ LTESGICFKS ERANGKLALD LDHGFACYMT SVGSPAECRF YSFRWKENEV LPSTVYA AN ITVSGHPGQP DLHFDSPEMI EFQSKKTGLM HYAMILRPSN FDPYKKYPVF HYVYGGPGIQ IVHNDFSWIQ YIRFCRLG Y VVVFIDNRGS AHRGIEFERH IHKKMGTVEV EDQVEGLQML AERTGGFMDM SRVVVHGWSY GGYMALQMIA KHPNIYRAA IAGGAVSDWR LYDTAYTERY MGYPLEEHVY GASSITGLVE KLPDEPNRLM LVHGLMDENV HFAHLTHLVD ECIKKGKWHE LVIFPNERH GVRNNDASIY LDARMMYFAQ QAIQGFGPTT AAPRQGPLWS HPQFEK UniProtKB: Dipeptidyl Peptidase Four (IV) family |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8pba: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)