+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 Spike RBD in complex with Mab-23 (Fab) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Antibody / Spike / Complex / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationMaturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell ...Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated endocytosis of virus by host cell / membrane fusion / Attachment and Entry / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / receptor ligand activity / symbiont-mediated suppression of host innate immune response / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Enterobacteria phage T4 (virus) / Enterobacteria phage T4 (virus) /  Homo sapiens (human) Homo sapiens (human) | |||||||||
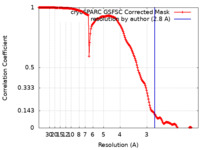
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Das H / Hallberg BM | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: SARS-CoV-2 Spike RBD in complex with Mab-23 (Fab domains) Authors: Das H / Hallberg BM | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17451.map.gz emd_17451.map.gz | 295.9 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17451-v30.xml emd-17451-v30.xml emd-17451.xml emd-17451.xml | 18.8 KB 18.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17451_fsc.xml emd_17451_fsc.xml | 16.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_17451.png emd_17451.png | 88.5 KB | ||
| Filedesc metadata |  emd-17451.cif.gz emd-17451.cif.gz | 6.2 KB | ||
| Others |  emd_17451_half_map_1.map.gz emd_17451_half_map_1.map.gz emd_17451_half_map_2.map.gz emd_17451_half_map_2.map.gz | 475.6 MB 475.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17451 http://ftp.pdbj.org/pub/emdb/structures/EMD-17451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17451 | HTTPS FTP |
-Validation report
| Summary document |  emd_17451_validation.pdf.gz emd_17451_validation.pdf.gz | 970.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_17451_full_validation.pdf.gz emd_17451_full_validation.pdf.gz | 969.7 KB | Display | |
| Data in XML |  emd_17451_validation.xml.gz emd_17451_validation.xml.gz | 23.5 KB | Display | |
| Data in CIF |  emd_17451_validation.cif.gz emd_17451_validation.cif.gz | 29.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17451 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17451 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17451 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17451 | HTTPS FTP |
-Related structure data
| Related structure data |  8p5mMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17451.map.gz / Format: CCP4 / Size: 1.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17451.map.gz / Format: CCP4 / Size: 1.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.01 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_17451_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17451_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Spike RBD (two molecules) in complex with two FU2 nanobodies
| Entire | Name: Spike RBD (two molecules) in complex with two FU2 nanobodies |
|---|---|
| Components |
|
-Supramolecule #1: Spike RBD (two molecules) in complex with two FU2 nanobodies
| Supramolecule | Name: Spike RBD (two molecules) in complex with two FU2 nanobodies type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Molecular weight | Theoretical: 95.7 KDa |
-Supramolecule #2: SARS-CoV-2 spike (RBD)
| Supramolecule | Name: SARS-CoV-2 spike (RBD) / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Monoclonal Antibody Mab-23
| Supramolecule | Name: Monoclonal Antibody Mab-23 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) |
-Macromolecule #1: Spike protein S1
| Macromolecule | Name: Spike protein S1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.002676 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: TNLCPFGEVF NATRFASVYA WNRKRISNCV ADYSVLYNSA SFSTFKCYGV SPTKLNDLCF TNVYADSFVI RGDEVRQIAP GQTGKIADY NYKLPDDFTG CVIAWNSNNL DSKVGGNYNY LYRLFRKSNL KPFERDISTE IYQAGSTPCN GVEGFNCYFP L QSYGFQPT ...String: TNLCPFGEVF NATRFASVYA WNRKRISNCV ADYSVLYNSA SFSTFKCYGV SPTKLNDLCF TNVYADSFVI RGDEVRQIAP GQTGKIADY NYKLPDDFTG CVIAWNSNNL DSKVGGNYNY LYRLFRKSNL KPFERDISTE IYQAGSTPCN GVEGFNCYFP L QSYGFQPT NGVGYQPYRV VVLSFELLHA PATVCGPK UniProtKB: Spike glycoprotein |
-Macromolecule #2: Mab-23 (Heavy chain variable domain)
| Macromolecule | Name: Mab-23 (Heavy chain variable domain) / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 48.500398 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLVESGGG LVQPGGSLRL SCTASGFTFS NYGFHWVRQA PGKGLEWVTI ISYDGITKHY ADSVKDRFTV SRDNSKTMVY LQMNNLKLD DTAVYYCARD LGTYDDSWGQ GVLVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGLVKDYF PEVTSWNSGA L TSGVHTFP ...String: EVQLVESGGG LVQPGGSLRL SCTASGFTFS NYGFHWVRQA PGKGLEWVTI ISYDGITKHY ADSVKDRFTV SRDNSKTMVY LQMNNLKLD DTAVYYCARD LGTYDDSWGQ GVLVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGLVKDYF PEVTSWNSGA L TSGVHTFP AVLQSSGLYS LSSVVTVPSS SLGTQTYICN VNHKPSNTKV DKVEPKSCDK THTCPPCPAP ELLGGPSVFL FP PKPKDTL MISRTPEVTC VVVDVSHEDP EVKFNWYVDG VEVHNAKTKP REEQYNSTYR VVSVLTVLHQ DWLNGKEYKC KVS NKALPA PIEKTISKAK GQPREPQVYT LPPSRDETKN QVSLTCLVKG FYPSDIAVEW ESNGQPENNY KTTPPVLDSD GSFF LYSKL TVDKSRWQQG NVFSCSVMHE ALHNHYTQKS LSLSPGK |
-Macromolecule #3: Mab-23 (Light chain variable domain)
| Macromolecule | Name: Mab-23 (Light chain variable domain) / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.234852 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIQMTQSPSA LSASVGDRVT ISCRASQNID GFLAWYQQKP GKAPKLLIYA ASRLQSGIPS RFSGSGSGTD FTLTISSLQP EDFAAYYCQ QVYSAPLTFG GGTKVEFKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS ...String: DIQMTQSPSA LSASVGDRVT ISCRASQNID GFLAWYQQKP GKAPKLLIYA ASRLQSGIPS RFSGSGSGTD FTLTISSLQP EDFAAYYCQ QVYSAPLTFG GGTKVEFKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS KDSTYSLSST LTLSKADYEK HKVYACEVTH QGLSSPVTKS FNRGEC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.1 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number real images: 4774 / Average exposure time: 1.5 sec. / Average electron dose: 48.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 20.0 µm / Calibrated defocus min: 0.3 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.3 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)