[English] 日本語
 Yorodumi
Yorodumi- EMDB-17003: Chaetomium thermophilum Methionine Aminopeptidase 2 at the 80S ri... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Chaetomium thermophilum Methionine Aminopeptidase 2 at the 80S ribosome | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ribosome Associated Factor / Protease / Tunnel exit / Protein Maturation / Proteostasis / NME / p67 / MAP / MetAP / MAP2 / MetAP2 / ES27L / PTE / METAL BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationdolichyl-diphosphooligosaccharide-protein glycotransferase / dolichyl-diphosphooligosaccharide-protein glycotransferase activity / protein N-linked glycosylation via asparagine / initiator methionyl aminopeptidase activity / methionyl aminopeptidase / protein kinase regulator activity / metalloaminopeptidase activity / protein-RNA complex assembly / cellular response to amino acid starvation / post-translational protein modification ...dolichyl-diphosphooligosaccharide-protein glycotransferase / dolichyl-diphosphooligosaccharide-protein glycotransferase activity / protein N-linked glycosylation via asparagine / initiator methionyl aminopeptidase activity / methionyl aminopeptidase / protein kinase regulator activity / metalloaminopeptidase activity / protein-RNA complex assembly / cellular response to amino acid starvation / post-translational protein modification / ribosomal large subunit biogenesis / regulation of translation / cytosolic large ribosomal subunit / cytoplasmic translation / rRNA binding / ribosome / structural constituent of ribosome / ribonucleoprotein complex / translation / proteolysis / RNA binding / membrane / metal ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Thermochaetoides thermophila (fungus) / Thermochaetoides thermophila (fungus) /  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.94 Å | |||||||||
 Authors Authors | Klein MA / Wild K / Kisonaite M / Sinning I | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Methionine aminopeptidase 2 and its autoproteolysis product have different binding sites on the ribosome. Authors: Marius A Klein / Klemens Wild / Miglė Kišonaitė / Irmgard Sinning /  Abstract: Excision of the initiator methionine is among the first co-translational processes that occur at the ribosome. While this crucial step in protein maturation is executed by two types of methionine ...Excision of the initiator methionine is among the first co-translational processes that occur at the ribosome. While this crucial step in protein maturation is executed by two types of methionine aminopeptidases in eukaryotes (MAP1 and MAP2), additional roles in disease and translational regulation have drawn more attention to MAP2. Here, we report several cryo-EM structures of human and fungal MAP2 at the 80S ribosome. Irrespective of nascent chains, MAP2 can occupy the tunnel exit. On nascent chain displaying ribosomes, the MAP2-80S interaction is highly dynamic and the MAP2-specific N-terminal extension engages in stabilizing interactions with the long rRNA expansion segment ES27L. Loss of this extension by autoproteolytic cleavage impedes interactions at the tunnel, while promoting MAP2 to enter the ribosomal A-site, where it engages with crucial functional centers of translation. These findings reveal that proteolytic remodeling of MAP2 severely affects ribosome binding, and set the stage for targeted functional studies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17003.map.gz emd_17003.map.gz | 1.3 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17003-v30.xml emd-17003-v30.xml emd-17003.xml emd-17003.xml | 28.3 KB 28.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_17003.png emd_17003.png | 66.4 KB | ||
| Filedesc metadata |  emd-17003.cif.gz emd-17003.cif.gz | 10.2 KB | ||
| Others |  emd_17003_half_map_1.map.gz emd_17003_half_map_1.map.gz emd_17003_half_map_2.map.gz emd_17003_half_map_2.map.gz | 1.3 GB 1.3 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17003 http://ftp.pdbj.org/pub/emdb/structures/EMD-17003 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17003 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17003 | HTTPS FTP |
-Validation report
| Summary document |  emd_17003_validation.pdf.gz emd_17003_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_17003_full_validation.pdf.gz emd_17003_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_17003_validation.xml.gz emd_17003_validation.xml.gz | 23.3 KB | Display | |
| Data in CIF |  emd_17003_validation.cif.gz emd_17003_validation.cif.gz | 28.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17003 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17003 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17003 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17003 | HTTPS FTP |
-Related structure data
| Related structure data |  8onzMC  8onxC  8onyC  8oo0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17003.map.gz / Format: CCP4 / Size: 1.4 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17003.map.gz / Format: CCP4 / Size: 1.4 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.868 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_17003_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
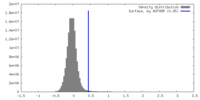
| Density Histograms |
-Half map: #1
| File | emd_17003_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
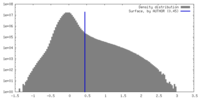
| Density Histograms |
- Sample components
Sample components
-Entire : MAP2 at the 80S ribosome
| Entire | Name: MAP2 at the 80S ribosome |
|---|---|
| Components |
|
-Supramolecule #1: MAP2 at the 80S ribosome
| Supramolecule | Name: MAP2 at the 80S ribosome / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#8 |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila (fungus) Thermochaetoides thermophila (fungus) |
-Macromolecule #1: Ribosomal protein L19
| Macromolecule | Name: Ribosomal protein L19 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Molecular weight | Theoretical: 317.092281 KDa |
| Recombinant expression | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Sequence | String: MSEPVANDAA PGSFDLEQAR TALTSSSTTV RIAQLRAIDE KISHNSLDRP ALLGLLKVLF WTHTFYADRP SRRAVQKCLT SITKCSEAE SFVPPLVAAI RQEAQKQAIA QANSFALVEW CSLLIQNLAG TSLWEKCSKD LVLATADALE KSLQPTARGS V GHSALVIT ...String: MSEPVANDAA PGSFDLEQAR TALTSSSTTV RIAQLRAIDE KISHNSLDRP ALLGLLKVLF WTHTFYADRP SRRAVQKCLT SITKCSEAE SFVPPLVAAI RQEAQKQAIA QANSFALVEW CSLLIQNLAG TSLWEKCSKD LVLATADALE KSLQPTARGS V GHSALVIT RRGFRKLVQA DQAAVSTAIG VLAAKDAKPT AKNSVLLGVI AGVCARQPEA KAVVEDLKGQ YFTFYAREII GS RTALPSH IADGLHDFFA AFVSLEDLDK EVFPAIEKGL LRAPEVVLND LITPLVKALP KLDLSKALNS RFVKPLLSNI KSS NAVIRS GAVRAFREIA TVCRDFADLE KTADEVLGPL KTNKLASADH RVLHSEMLSG LPLSASIATK IATGLPAVTS KEAN EAALS AETLALNASA LFLLRTPEVP KALLDAYVKG LGDKKVSSRR IWILRTGDLL QAFAQEAELP AGFVAFAEAV VPSLL TIFN EVVANPTAAV QSGLVTGALV VAGLSGLLQR LENANIQALL KKAAVQKNSL ILDPKPSFVL SQRLYSKYPE DDLKWF CRA LSAIVEGLLS SSDAIKLAWT QAFIFLICSA TTPSAVRRQA IDALSDLSAR SLKPGCQFVA TEIIKGLWHW IATSEAA EK ESAAVLAKSG TSNLHLVLKA ICLSPAELNR RSGGDVDKAQ LQAQMCSLLV LAKQQLIPRA SWIDLCLKVQ VDPGQLAK D HEDALIDEIV RCTGHEQKSE ATKQAAYSAA AELVFVSPEN MTSRIVALIQ DDLDAAVVST VGPLEAAIFR TPEGTTFVD VLAKRQNVVP NKNVKDYDLL KWEEELRQQL AEKKGIQKKL TAEEQAKVSA QLKKEAEIRE NVRKIAAKLL RGFGVVKALA TGPPTDATR WMGAAVSSTL AAINAGATRI TGETGPLAFV SLSEQITSRI GSFIRPFMGV ATLRAYEVTA LPENLTEEPF E ELITRVLY RLRFVGEQRP LDTVSLIYVL PLVLYVLEKG GFAKDADEKD AQLVLAIEIL SFHTDVAFDE ALPRAEILSS LI SSMQKYS QHYKILKDCF SDMVRCIAPN ISPAEIGVLA RGAIVPQTSV RTAVLQAISA DVDMSDVNAS EEIWLACHDD IDE NAELGR EIWEESEWKT SEELGHKMIP YLESKDVQLR RAAAKSLAEV AGQHPDVVAP ILEKLRESYV ELAKPRVQQL DEFG MPKKM DLSDPWEARH GIALAFKGLA PHLEKRQLEP YFNFLIEQGP LGDQSAGVRA EMLEAANMTI EIHGKEILDR LMKTF EKVL EAPDKNSEAA DRVNEAVIIM YGALARHLKP GDKKIPVVIE RLLATLSTPS EAVQYAIAEC LPPLVRTCAD KSSKYF DEM MEILLTSKKY SEQRGAAYGL AGLVLGRGIN VLKEYRIMTQ LNSALENKKE IRQRESAMIA YELLSTILGR LFEPYVI QI VPQLLAGFGD GNADVREAAL AAAKACFAKL SSYGVKQILP TLLNGLDDDQ WRSKKGACDL LGAMAYLDPQ QLAQNLPE I IPPLTAVLND SHKEVRAAAN RSLKRFGEVI TNPEIKSLID ILLKALSDPT KYTDDALDAL IKVQFVHYLD APSLALVSR ILQRGLGDRS NTKRKASQVI GSLAHLTERK DLIAHLPVLV AGLKVAVVDP VPTTRATASR ALGSLVEKLG EDALPDLIPN LMQTLKSDT SAGDRLGSAQ ALSEVLAGLG TSRLEETLPT ILQNVESPKP AVREGFMSLF IFLPVCFGNS FANYLGKIIP P ILSGLADD VESIRETALR AGRLLVKNFA VRAVDLLLPE LERGLADDNY RIRLSSVELV GDLLFNLAGV KASANKEEDE AD QDITKEA GASLREVLGE EKRNKILSAL YVCRCDTAGA VRAAAVTVWK QLVHSPRTLK ELVPTLTQLL IKRLGSSNME HKV IASNAL GELIKKAGDG VLATLLPTLE EGLQTSRDVN AKQGICLALK ELISSASPEA LEDHEKTLIS VVRTALTDSD SDVR EAAAE AFDSLQQIIG KRAIDQVLPF LLNLLRSEEE ANNALAALLT LLTEPTRANI ILPNLIPTLI TPPISAFNAK ALASL SKVA GAAMNRRLPN IINSLMDNIV GCADETLREE LDTSFDTVIL SIDDTDGLNV VMNVLLQLIK HEDHRKRAAT GRHLAK FFS AATVNYSRYN QDIIRALLIS FDDKDMEVVK ASWNALNEFT KRLKKEEMEG LVISTRQTLL QVGVAGRELA GFELPKG IN AILPIFLQGL MNGTAEQRVA AALGISDIVD RTTEASLKPF VTQITGPLIR VVSERSTEVR SAILLTLNHL LEKMPTAL K PFLPQLQRTF AKSLADTSSE ILRSRAARAL GTLIKYTPRV DPLIAELVTG SKTTDAGVKT AMLKALYEVI SKAGSNMGE SSRAAVLGLI DMEPDENDKA MTITNAKLFG ALMKNVPADM AVGLLKNRVM VREFTTSSVL ALNAVLLESP SSLLDSPLAE DLPELLCQG IESKDAFIAD NFIMATGKYL LHDSPKAFET TKAIFTSLAK LIPPGNPGDS RRLSLVLVRT MARKQPDMVR P HLSLLAPP IFSSVRDMVI PVKLAAEAAF VQLFAVADEE SKVFDKWITA ATDLAPNVKR SMQDYFKRVT LRLGAQARER RE AEGGAGT LGLSADEVED EKELMAVGKP RVSWNRNPEK LKKPRAQSEV DPGPVPLKTV ARARQKSFKM VNLRTQKRLA ASV LGCGEG KVWLDPNEVS EISNANSRQS IRKLVADGLI IKKPVTMHSR SRARELNLAR RIGRHRGFGK RKGTADARMP QQVL WMRRQ RVLRRLLVKY RASGKIDKHL YHELYHLAKG NTFKHKRALV EHIHRAKAEK ARERQIKEEM DAKRARTKAA RERKQ ERQA AKRNALLGEE EESK UniProtKB: Ribosomal protein L19 |
-Macromolecule #2: 60S ribosomal protein L25-like protein
| Macromolecule | Name: 60S ribosomal protein L25-like protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Molecular weight | Theoretical: 17.175334 KDa |
| Recombinant expression | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Sequence | String: MAPKDVKKGG ASKAAKGAQA KKAAQAALKG VHSHKKTKVR KSTTFHRPKT LVLSRAPKYP RKSIPHEPRL DEHKIIIHPL NTEGALKKI EEQNTLVFIV DVKANKAQIK QALKKLYDID TVKINTLIRP DGTKKAFARL TPDVDALDIA ATKLGLV UniProtKB: 60S ribosomal protein L25-like protein |
-Macromolecule #3: 60S ribosomal protein L26-like protein
| Macromolecule | Name: 60S ribosomal protein L26-like protein / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Molecular weight | Theoretical: 15.565322 KDa |
| Recombinant expression | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Sequence | String: MKVRPTVSSS RRKARKAHFS APSSVRRVIM SAPLSKELRE KYNVRSIPIR KGDEVQIVRG AHKDKEGKVT SVYRLKYVIH VERVTREKA TGQTVPIGIH PSNVVITKLH LDKDRENILA RIKAGREQVA KAKGKKTAA UniProtKB: 60S ribosomal protein L26-like protein |
-Macromolecule #4: dolichyl-diphosphooligosaccharide--protein glycotransferase
| Macromolecule | Name: dolichyl-diphosphooligosaccharide--protein glycotransferase type: protein_or_peptide / ID: 4 Details: This protein is wrongly annoted as dolichyl-diphosphooligosaccharide-protein glycotransferase in uniprotKB, which explains the long non-matching C-terminus. Number of copies: 1 / Enantiomer: LEVO EC number: dolichyl-diphosphooligosaccharide-protein glycotransferase |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Molecular weight | Theoretical: 105.169227 KDa |
| Recombinant expression | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Sequence | String: MSSNGKVKAG QLWSKNKEEL TKILGELKTE LSQLRIQKIS SSGAKLNKIH DLRKSIARVL TVINAKQRAQ LRLFYKNKKY LPLDLRPKL TRALRRRLSK EDASRVLEKT KKRLTHFPQR KYAVKAATYS EGPALSIYHS HQREARCSLV QCCTFEPELR E SLLLLPVA ...String: MSSNGKVKAG QLWSKNKEEL TKILGELKTE LSQLRIQKIS SSGAKLNKIH DLRKSIARVL TVINAKQRAQ LRLFYKNKKY LPLDLRPKL TRALRRRLSK EDASRVLEKT KKRLTHFPQR KYAVKAATYS EGPALSIYHS HQREARCSLV QCCTFEPELR E SLLLLPVA ASLKGPNFTR RERQHRNPSE ATMSAAEPLK LLASAGKGKS TRSVLRVAIL VLIAGAAVAS RLFSVIRFES II HEFDPWF NFRATKYLVA NGFYKFWDWF DDRTWHPLGR VTGGTLYPGL MVTSGVIYHL LRFLTVPVDI RNICVLLAPG FSG LTAIAA YLLTNEMTTS PSAGLLAAAF MGIAPGYISR SVAGSYDNEA IAIFLLVFTF FLWIKALKQG SMLWGALCAL FYGY MVASW GGYAFITCLL PLHSFVLICM GRYSTRLYVA YTTWYALGTL ASMQIPFVGF LPVKTSEHMP ALGIFGFLQL LAFLD YVRS TISSRQFQTF LWLFAGGIFG LGLLGLVIAT SAGLIAPWSG RFYSLWDTGY AKIHIPIIAS VSEHQPTAWP AFFFDL NML VWLFPVGVYL CFQQLGDEHV FIIVYALFGS YFAGVMVRLM LTLTPVVCVA AAIAFSSLLD TYLNLKTPNP GQAQATE DA GKKKSGLKAA SKPAVGVYAL WGKTMMISGL TIYLLLFVLH CTWVTSNAYS SPSVVLASRL PDGSQHIIDD YREAYQWL R QNTREDAKIM SWWDYGYQIG GMADRPTLVD NNTWNNTHIA TVGKAMASRE EVSYPIMRQH EVDYVLVVFG GLLGYSGDD INKFLWMVRI AEGIWPDEVS ERAFFTPRGE YRVDAEATDT MKNSLMYKMC YYNYNNLFPP GQAVDRMRGV RLPEVGPTLN TLEEAFTSE NWIIRIYKVK DLDNLGRDHA SAAAFERGHK KKKATKKRGP RVLRVE UniProtKB: dolichyl-diphosphooligosaccharide--protein glycotransferase |
-Macromolecule #5: 60S ribosomal protein L38-like protein
| Macromolecule | Name: 60S ribosomal protein L38-like protein / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Molecular weight | Theoretical: 10.902031 KDa |
| Recombinant expression | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Sequence | String: MPQEVSDIKK FIEICRRKDA SSKILTIAFP PPLTAARIKK NPKTQQIKFK VRCQRFLYTL VLKDSDKAEK LKQSLPPNLQ IKDVPKRNK RKSSA UniProtKB: 60S ribosomal protein L38-like protein |
-Macromolecule #8: Methionine aminopeptidase 2
| Macromolecule | Name: Methionine aminopeptidase 2 / type: protein_or_peptide / ID: 8 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Molecular weight | Theoretical: 48.912805 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAAQVPTEEL SKLSVQDAAA VKPGADAPEP ANGKENESDD EEDEAEEAAA APESGAGGAK KKKKKKNKKK KKKPTQQSDP PRVLISHLF PDGKYPAGEE VEYVNENRYR TTSEEKRYLD NMQSEFLNDY RQAAEVHRQV RKWAQGFVKP GKSLIEISEG I EDSVRALV ...String: MAAQVPTEEL SKLSVQDAAA VKPGADAPEP ANGKENESDD EEDEAEEAAA APESGAGGAK KKKKKKNKKK KKKPTQQSDP PRVLISHLF PDGKYPAGEE VEYVNENRYR TTSEEKRYLD NMQSEFLNDY RQAAEVHRQV RKWAQGFVKP GKSLIEISEG I EDSVRALV GHPGLEEGDA LKAGMGFPVG LSINHCAAHY NPNSGNKIVL QQDDVIKIDI GVHVNGRIVD SAFTMAWNDQ FN PLLEAVR AATNAGIREA GIDARVGEIG GVIQEVMESY EVEINGKTYP VKPIRNLNGH NILPYSIHGT KSVPIVKTHD QTK MEEGDV FAIETFGSTG KGYVIESGEV SHYALRGDAP KVDLRLSSAK SLLNVIKRHF GTLPFCRRFL DRLGQEKYLL GLNN LVSNG IVEDYPPLVD EKGSYTAQFE HTILIRPTVK EVISRGDDY UniProtKB: Methionine aminopeptidase 2 |
-Macromolecule #6: 28S rRNA
| Macromolecule | Name: 28S rRNA / type: rna / ID: 6 Details: alignment does not work properly here in context of 28S rRNA sequence ... to be redone Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Molecular weight | Theoretical: 1.078716125 MDa |
| Sequence | String: AGGUUGACCU CGGAUCAGGU AGGAGGACCC GCUGAACUUA AGCAUAUCAA UAAGCGGAGG AAAAGAAACC AACAGGGAUU GCCCUAGUA ACGGCGAGUG AAGCGGCAAC AGCUCAAAUU UGAAAGCUGG CUUCGGCCCG CGUUGUAAUU UGGAGAGGAU G CUUUGGGC ...String: AGGUUGACCU CGGAUCAGGU AGGAGGACCC GCUGAACUUA AGCAUAUCAA UAAGCGGAGG AAAAGAAACC AACAGGGAUU GCCCUAGUA ACGGCGAGUG AAGCGGCAAC AGCUCAAAUU UGAAAGCUGG CUUCGGCCCG CGUUGUAAUU UGGAGAGGAU G CUUUGGGC GAGGCUCCUU CUGAGUUCCC UGGAACGGGA CGCCACAGAG GGUGAGAGCC CCGUAUAGUU GGAAGCCAAG CC UGUGUAA AGCUCCUUCG ACGAGUCGAG UAGUUUGGGA AUGCUGCUCA AAAUGGGAGG UAAAUUUCUU CUAAAGCUAA AUA CCGGCC AGAGACCGAU AGCGCACAAG UAGAGUGAUC GAAAGAUGAA AAGCACUUUG AAAAGAGGGU UAAAUAGCAC GUGA AAUUG UUGAAAGGGA AGCGCUUGUG ACCAGACUUG CGCCCGGCGG AUCAUCCGGU GUUCUCACCG GUGCACUCCG CCGGG CUCA GGCCAGCAUC GGUUCUGGCG GGGGGAUAAA GGCCCAGGGA AUGUGGCUCC UCCGGGAGUG UUAUAGCCCU GGGUGU AAU ACCCUCGCCG GGACCGAGGA CCGCGCUCUG CAAGGAUGCU GGCGUAAUGG UCACCAGCGA CCCGUCUUGA AACACGG AC CAAGGAGUCA AGGUUUUGCG CGAGUGUUUG GGUGUAAAAC CCGCACGCGU AAUGAAAGUG AACGUAGGUG AGAGCUUC G GCGCAUCAUC GACCGAUCCU GAUGUAUUCG GAUGGAUUUG AGUAGGAGCG UUAAGCCUUG GACCCGAAAG AUGGUGAAC UAUGCUUGGA UAGGGUGAAG CCAGAGGAAA CUCUGGUGGA GGCUCGCAGC GGUUCUGACG UGCAAAUCGA UCGUCAAAUC UGAGCAUGG GGGCGAAAGA CUAAUCGAAC CAUCUAGUAG CUGGUUACCG CCGAAGUUUC CCUCAGGAUA GCAGUGUCGA C CUUCAGUU UUAUGAGGUA AAGCGAAUGA UUAGGGACUC GGGGGCGAUU UUUAGCCUUC AUCCAUUCUC AAACUUUAAA UA UGUAAGA AGCCCUUGUU ACUUAACUGA ACGUGGGCAU UCGAAUGUAU CGACACUAGU GGGCCAUUUU UGGUAAGCAG AAC UGGCGA UGCGGGAUGA ACCGAACGCG GGGUUAAGGU GCCGGAGUGG ACGCUCAUCA GACACCACAA AAGGCGUUAG UACA UCUUG ACAGCAGGAC GGUGGCCAUG GAAGUCGGAA UCCGCUAAGG ACUGUGUAAC AACUCACCUG CCGAAUGUAC UAGCC CUGA AAAUGGAUGG CGCUCAAGCG UCCCACCCAU ACCCCGCCCU CAGGGUAGAA ACGAUGCCCU GAGGAGUAGG CGGCCG UGG AGGUCAGUGA CGAAGCCUAG GGCGUGAGCC CGGGUCGAAC GGCCUCUAGU GCAGAUCUUG GUGGUAGUAG CAAAUAC UU CAAUGAGAAC UUGAAGGACC GAAGUGGGGA AAGGUUCCAU GUGAACAGCG GUUGGACAUG GGUUAGUCGA UCCUAAGC C AUAGGGAAGU UCCGUUUCAA AGGGGCACUC GUGCCCCGUG UGGCGAAAGG GAAGCCGGUU AAUAUUCCGG CACCUGGAU GUGGGUUUUG CGCGGCAACG CAACUGAACG CGGAGACGAC GGCGGGGGCC CCGGGCAGAG UUCUCUUUUC UUCUUAACGG UCUAUCACC CUGGAAACAG UUUGUCUGGA GAUAGGGUUU AAUGGCCGGA AGAGCCCGAC ACUUCUGUCG GGUCCGGUGC G CUCUCGAC GUCCCUUGAA AAUCCGCGGG AGGGAAUAAU UCUCACGCCA GGUCGUACUC AUAACCGCAG CAGGUCCCCA AG GUGAACA GCCUCUGGUU GAUAGAACAA UGUAGAUAAG GGAAGUCGGC AAAAUAGAUC CGUAACUUCG GGAAAAGGAU UGG CUCUAA GGGUUGGGCA CGUUGGGCUU UGGGCGGACG CCCUGGGAGC AGAGGGCCUC UAGCCGGGCA ACCGGCCGGC GGCC CUCAG CACCCGGGGU UGAAGCCCUU AGCAGGCUUC GGCCGUCCGG CGUGCGGUUA ACAACCAACU UAGAACUGGU ACGGA CAGG GGGAAUCUGA CUGUCUAAUU AAAACAUAGC AUUGCGAUGG CCAGAAAGUG GUGUUGACGC AAUGUGAUUU CUGCCC AGU GCUCUGAAUG UCAAAGUGAA GAAAUUCAAC CAAGCGCGGG UAAACGGCGG GAGUAACUAU GACUCUCUUA AGGUAGC CA AAUGCCUCGU CAUCUAAUUA GUGACGCGCA UGAAUGGAUU AACGAGAUUC CCACUGUCCC UAUCUACUAU CUAGCGAA A CCACAGCCAA GGGAACGGGC UUGGCAAAAU CAGCGGGGAA AGAAGACCCU GUUGAGCUUG ACUCUAGUUU GACAUUGUG AAAAGACAUA GGAGGUGUAG AAUAGGUGGG AGCUUCGGCG CCAGUGAAAU ACCACUACUC CUAUUGUUUU UUUACUUAUU CAAUGAAGC GGGGCUGGAC UUGCGUCCAA CUUCUGGAGU UAAGGUCCUU CGCGGGCCGA CCCGGGUUGA AGACAUUGUC A GGUGGGGA GUUUGGCUGG GGCGGCACAU CUGUUAAACC AUAACGCAGG UGUCCUAAGG GGGGCUCAUG GAGAACAGAA AU CUCCAGU AGAACAAAAG GGUAAAAGUC CCCUUGAUUU UGAUUUUCAG UGUGAAUACA AACCAUGAAA GUGUGGCCUA UCG AUCCUU UAGUCCCUCG AAAUUUGAGG CUAGAGGUGC CAGAAAAGUU ACCACAGGGA UAACUGGCUU GUGGCGGCCA AGCG UUCAU AGCGACGUCG CUUUUUGAUC CUUCGAUGUC GGCUCUUCCU AUCAUACCGA AGCAGAAUUC GGUAAGCGUU GGAUU GUUC ACCCACUAAU AGGGAACGUG AGCUGGGUUU AGACCGUCGU GAGACAGGUU AGUUUUACCC UACUGAUGAA CUCGUC GCA AUGGUAAUUC AGCUUAGUAC GAGAGGAACC GCUGAUUCAG AUAAUUGGUU UUUGCGGUUG UCCGACCGGG CAGUGCC GC GAAGCUACCA UCUGCUGGAU AAUGGCUGAA CGCCUCUAAG UCAGAAUCCA UGCCAGAACG CGACGAUACU ACCCGCAC G UUGUAGACGU AUAAGAAUAG GCUCCGGCCU CGUAUCCUAG CAGGCGAUUC CUCCGCCGGC CUCGAAGUGG CCGUCGGUA AUUCGCGUAU UGCAAUUUAG ACACGCGCGG GAUCAAAUCC UUUGCAGACG ACUUAGAUGU GCGAAAGGGU CCUGUAAGCA GUAGAGUAG CCUUGUUGUU ACGAUCUGCU GAGGGUAAGC CCUCCUUCGC CUAGAUUU GENBANK: GENBANK: XR_002966752.1 |
-Macromolecule #7: 5.8S rRNA
| Macromolecule | Name: 5.8S rRNA / type: rna / ID: 7 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila DSM 1495 (fungus) Thermochaetoides thermophila DSM 1495 (fungus) |
| Molecular weight | Theoretical: 50.147688 KDa |
| Sequence | String: AAACUUUCAA CAACGGAUCU CUUGGUUCUG GCAUCGAUGA AGAACGCAGC GAAAUGCGAU AAGUAAUGUG AAUUGCAGAA UUCCGUGAA UCAUCGAAUC UUUGAACGCA CAUUGCGCCC GCCGGUAUUC CGGCGGGCAU GCCUGUUCGA GCGUCAU GENBANK: GENBANK: XR_002966750.1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 42.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.1 µm / Nominal defocus min: 1.1 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.94 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 61734 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)