+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
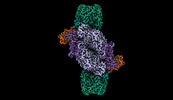
| Title | Iron Nitrogenase Complex from Rhodobacter capsulatus | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | nitrogen fixation / Fe nitrogenase / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationnitrogenase / : / nitrogenase activity / nitrogen fixation / iron-sulfur cluster binding / 4 iron, 4 sulfur cluster binding / iron ion binding / ATP binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Rhodobacter capsulatus SB 1003 (bacteria) Rhodobacter capsulatus SB 1003 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.35 Å | |||||||||
 Authors Authors | Schmidt FV / Schulz L / Zarzycki J / Prinz S / Erb TJ / Rebelein JG | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Structural insights into the iron nitrogenase complex. Authors: Frederik V Schmidt / Luca Schulz / Jan Zarzycki / Simone Prinz / Niels N Oehlmann / Tobias J Erb / Johannes G Rebelein /  Abstract: Nitrogenases are best known for catalyzing the reduction of dinitrogen to ammonia at a complex metallic cofactor. Recently, nitrogenases were shown to reduce carbon dioxide (CO) and carbon monoxide ...Nitrogenases are best known for catalyzing the reduction of dinitrogen to ammonia at a complex metallic cofactor. Recently, nitrogenases were shown to reduce carbon dioxide (CO) and carbon monoxide to hydrocarbons, offering a pathway to recycle carbon waste into hydrocarbon products. Among the three nitrogenase isozymes, the iron nitrogenase has the highest wild-type activity for the reduction of CO, but the molecular architecture facilitating these activities has remained unknown. Here, we report a 2.35-Å cryogenic electron microscopy structure of the ADP·AlF-stabilized iron nitrogenase complex from Rhodobacter capsulatus, revealing an [FeSC-(R)-homocitrate] cluster in the active site. The enzyme complex suggests that the iron nitrogenase G subunit is involved in cluster stabilization and substrate channeling and confers specificity between nitrogenase reductase and catalytic component proteins. Moreover, the structure highlights a different interface between the two catalytic halves of the iron and the molybdenum nitrogenase, potentially influencing the intrasubunit 'communication' and thus the nitrogenase mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16890.map.gz emd_16890.map.gz | 108 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16890-v30.xml emd-16890-v30.xml emd-16890.xml emd-16890.xml | 22.4 KB 22.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16890_fsc.xml emd_16890_fsc.xml | 12.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_16890.png emd_16890.png | 45.1 KB | ||
| Filedesc metadata |  emd-16890.cif.gz emd-16890.cif.gz | 6.9 KB | ||
| Others |  emd_16890_half_map_1.map.gz emd_16890_half_map_1.map.gz emd_16890_half_map_2.map.gz emd_16890_half_map_2.map.gz | 193.8 MB 193.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16890 http://ftp.pdbj.org/pub/emdb/structures/EMD-16890 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16890 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16890 | HTTPS FTP |
-Related structure data
| Related structure data |  8oieMC  8pbbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16890.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16890.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.837 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_16890_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_16890_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Iron Nitrogenase Complex from Rhodobacter capsulatus
+Supramolecule #1: Iron Nitrogenase Complex from Rhodobacter capsulatus
+Macromolecule #1: Nitrogenase protein alpha chain
+Macromolecule #2: Nitrogenase iron-iron protein delta chain
+Macromolecule #3: Nitrogenase iron-iron protein, beta subunit
+Macromolecule #4: Nitrogenase iron protein
+Macromolecule #5: FeFe cofactor
+Macromolecule #6: 3-HYDROXY-3-CARBOXY-ADIPIC ACID
+Macromolecule #7: FE(8)-S(7) CLUSTER
+Macromolecule #8: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #9: MAGNESIUM ION
+Macromolecule #10: ALUMINUM FLUORIDE
+Macromolecule #11: IRON/SULFUR CLUSTER
+Macromolecule #12: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 Details: 50 mM TRIS (pH = 7.8) 200 mM NaCl 5 mM sodium dithionite |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X