+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of PcrV/Fab(11-E5) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | antibody T3SS Tip protein / ANTIMICROBIAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationtype III protein secretion system complex / protein secretion by the type III secretion system / detection of maltose stimulus / maltose transport complex / carbohydrate transport / carbohydrate transmembrane transporter activity / maltose binding / maltose transport / maltodextrin transmembrane transport / ATP-binding cassette (ABC) transporter complex, substrate-binding subunit-containing ...type III protein secretion system complex / protein secretion by the type III secretion system / detection of maltose stimulus / maltose transport complex / carbohydrate transport / carbohydrate transmembrane transporter activity / maltose binding / maltose transport / maltodextrin transmembrane transport / ATP-binding cassette (ABC) transporter complex, substrate-binding subunit-containing / ATP-binding cassette (ABC) transporter complex / cell chemotaxis / outer membrane-bounded periplasmic space / periplasmic space / DNA damage response / extracellular space / membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
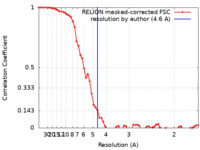
| Method | single particle reconstruction / cryo EM / Resolution: 4.6 Å | |||||||||
 Authors Authors | Yuan B / Simonis A / Marlovits TC | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2023 Journal: Cell / Year: 2023Title: Discovery of highly neutralizing human antibodies targeting Pseudomonas aeruginosa. Authors: Alexander Simonis / Christoph Kreer / Alexandra Albus / Katharina Rox / Biao Yuan / Dmitriy Holzmann / Joana A Wilms / Sylvia Zuber / Lisa Kottege / Sandra Winter / Meike Meyer / Kristin ...Authors: Alexander Simonis / Christoph Kreer / Alexandra Albus / Katharina Rox / Biao Yuan / Dmitriy Holzmann / Joana A Wilms / Sylvia Zuber / Lisa Kottege / Sandra Winter / Meike Meyer / Kristin Schmitt / Henning Gruell / Sebastian J Theobald / Anna-Maria Hellmann / Christina Meyer / Meryem Seda Ercanoglu / Nina Cramer / Antje Munder / Michael Hallek / Gerd Fätkenheuer / Manuel Koch / Harald Seifert / Ernst Rietschel / Thomas C Marlovits / Silke van Koningsbruggen-Rietschel / Florian Klein / Jan Rybniker /  Abstract: Drug-resistant Pseudomonas aeruginosa (PA) poses an emerging threat to human health with urgent need for alternative therapeutic approaches. Here, we deciphered the B cell and antibody response to ...Drug-resistant Pseudomonas aeruginosa (PA) poses an emerging threat to human health with urgent need for alternative therapeutic approaches. Here, we deciphered the B cell and antibody response to the virulence-associated type III secretion system (T3SS) in a cohort of patients chronically infected with PA. Single-cell analytics revealed a diverse B cell receptor repertoire directed against the T3SS needle-tip protein PcrV, enabling the production of monoclonal antibodies (mAbs) abrogating T3SS-mediated cytotoxicity. Mechanistic studies involving cryoelectron microscopy identified a surface-exposed C-terminal PcrV epitope as the target of highly neutralizing mAbs with broad activity against drug-resistant PA isolates. These anti-PcrV mAbs were as effective as treatment with conventional antibiotics in vivo. Our study reveals that chronically infected patients represent a source of neutralizing antibodies, which can be exploited as therapeutics against PA. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16807.map.gz emd_16807.map.gz | 2.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16807-v30.xml emd-16807-v30.xml emd-16807.xml emd-16807.xml | 16.7 KB 16.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16807_fsc.xml emd_16807_fsc.xml | 6.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_16807.png emd_16807.png | 56.5 KB | ||
| Filedesc metadata |  emd-16807.cif.gz emd-16807.cif.gz | 6.2 KB | ||
| Others |  emd_16807_half_map_1.map.gz emd_16807_half_map_1.map.gz emd_16807_half_map_2.map.gz emd_16807_half_map_2.map.gz | 13.8 MB 13.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16807 http://ftp.pdbj.org/pub/emdb/structures/EMD-16807 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16807 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16807 | HTTPS FTP |
-Validation report
| Summary document |  emd_16807_validation.pdf.gz emd_16807_validation.pdf.gz | 772.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16807_full_validation.pdf.gz emd_16807_full_validation.pdf.gz | 771.7 KB | Display | |
| Data in XML |  emd_16807_validation.xml.gz emd_16807_validation.xml.gz | 11.5 KB | Display | |
| Data in CIF |  emd_16807_validation.cif.gz emd_16807_validation.cif.gz | 15.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16807 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16807 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16807 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16807 | HTTPS FTP |
-Related structure data
| Related structure data |  8crbMC  8cr9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16807.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16807.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_16807_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_16807_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PcrV-Fab(11-E5)
| Entire | Name: PcrV-Fab(11-E5) |
|---|---|
| Components |
|
-Supramolecule #1: PcrV-Fab(11-E5)
| Supramolecule | Name: PcrV-Fab(11-E5) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Heavy chain
| Macromolecule | Name: Heavy chain / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.036674 KDa |
| Sequence | String: EVQLLESGGG LVQPGGSLRL SCAASGFSFS SYAMSWVRQA PGKGLEWVSA ISGSGGITYY GDSAKGRFTI SRDNSKNTLY LEMSSLRAD DTAVYYCAQE RYCDSGSCYE RDPVFEYWGQ GTRVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY F PEPVTVSW ...String: EVQLLESGGG LVQPGGSLRL SCAASGFSFS SYAMSWVRQA PGKGLEWVSA ISGSGGITYY GDSAKGRFTI SRDNSKNTLY LEMSSLRAD DTAVYYCAQE RYCDSGSCYE RDPVFEYWGQ GTRVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY F PEPVTVSW NSGALTSGVH TFPAVLQSSG LYSLSSVVTV PSSSLGTQTY ICNVNHKPSN TKVDKRVEP |
-Macromolecule #2: Light chain
| Macromolecule | Name: Light chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 22.964422 KDa |
| Sequence | String: QSVLTQPPSA SGAPGQRVTI SCSGSNSNIG TYFVYWYQQL PGTAPKVLIY RNDQRPSGVP DRISGSKSGT SASLAISGLR SEDEADYYC ASWDASLRGY VFGPGTKVTV LGQPKAAPSV TLFPPSSEEL QANKATLVCL ISDFYPGAVT VAWKADSSPV K AGVETTTP ...String: QSVLTQPPSA SGAPGQRVTI SCSGSNSNIG TYFVYWYQQL PGTAPKVLIY RNDQRPSGVP DRISGSKSGT SASLAISGLR SEDEADYYC ASWDASLRGY VFGPGTKVTV LGQPKAAPSV TLFPPSSEEL QANKATLVCL ISDFYPGAVT VAWKADSSPV K AGVETTTP SKQSNNKYAA SSYLSLTPEQ WKSHRSYSCQ VTHEGSTVEK TVAPTECS |
-Macromolecule #3: Maltose/maltodextrin-binding periplasmic protein,Type III secreti...
| Macromolecule | Name: Maltose/maltodextrin-binding periplasmic protein,Type III secretion protein PcrV type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 75.538055 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSWSHPQFE KGGGSGGGSG GGSWSHPQFE KSGLVPRGSA SKTEEGKLVI WINGDKGYNG LAEVGKKFEK DTGIKVTVEH PDKLEEKFP QVAATGDGPD IIFWAHDRFG GYAQSGLLAE ITPAAAFQDK LYPFTWDAVR YNGKLIAYPI AVEALSLIYN K DLLPNPPK ...String: MGSWSHPQFE KGGGSGGGSG GGSWSHPQFE KSGLVPRGSA SKTEEGKLVI WINGDKGYNG LAEVGKKFEK DTGIKVTVEH PDKLEEKFP QVAATGDGPD IIFWAHDRFG GYAQSGLLAE ITPAAAFQDK LYPFTWDAVR YNGKLIAYPI AVEALSLIYN K DLLPNPPK TWEEIPALDK ELKAKGKSAL MFNLQEPYFT WPLIAADGGY AFKYAAGKYD IKDVGVDNAG AKAGLTFLVD LI KNKHMNA DTDYSIAEHA FNHGETAMTI NGPWAWSNID TSKVNYGVTV LPTFKGQPSK PFVGVLSAGI NAASPNKELA KEF LENYLL TDEGLEAVNK DKPLGAVALK SYEEELVKDP RVAATMENAQ KGEIMPNIPQ MSAFWYAVRT AVINAASGRQ TVDE VRNLN AARELFLDEL LAAPAAPASA EQEELLALLR SERIVLAHAG QPLSEAQVLK ALAWLLAANP SAPPGQGLEV LREVL QARR QPGAQWDLRE FLVSAYFSLH GRLDEDVIGV YKDVLQTQDG KRKALLDELK ALTAELKVYS VIQSQINAAL SAKQGI RID AGGIDLVDPT LYGYAVGDPR WKDSPEYALL SNLDTFSGKL SIKDFLSGSP KQSGELKGLK DEYPFEKDNN PVGNFAT TV SDRSRPLNDK VNEKTTLLND TSSRYNSAVE ALNRFIQKYD SVLRDILSAI UniProtKB: Maltose/maltodextrin-binding periplasmic protein, Type III secretion protein PcrV |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.33 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 1xPBS |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 1.24 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X