+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Carin1 bacteriophage portal assembly | |||||||||
 Map data Map data | ||||||||||
 Sample Sample | Bacteriophage sp. Cobetia Marina Virus Carin1 != Bacteriophage sp. Bacteriophage sp. Cobetia Marina Virus Carin1
| |||||||||
 Keywords Keywords | Bacteriophages / virus structure / Cryo-EM / marine podovirus / portal protein / tail / VIRUS | |||||||||
| Biological species |  Bacteriophage sp. (virus) Bacteriophage sp. (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | d'Acapito A / Neumann E / Schoehn G | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: J Virol / Year: 2023 Journal: J Virol / Year: 2023Title: Structural Study of the Cobetia marina Bacteriophage 1 (Carin-1) by Cryo-EM. Authors: Alessio d'Acapito / Thomas Roret / Eleftherios Zarkadas / Pierre-Yves Mocaër / Florian Lelchat / Anne-Claire Baudoux / Guy Schoehn / Emmanuelle Neumann /  Abstract: Most of studied bacteriophages (phages) are terrestrial viruses. However, marine phages are shown to be highly involved in all levels of oceanic regulation. They are, however, still largely ...Most of studied bacteriophages (phages) are terrestrial viruses. However, marine phages are shown to be highly involved in all levels of oceanic regulation. They are, however, still largely overlooked by the scientific community. By inducing cell lysis on half of the bacterial population daily, their role and influence on the bacterial biomass and evolution, as well as their impact in the global biogeochemical cycles, is undeniable. Cobetia marina (Carin-1) is a member of the family infecting the γ. marina. Here, we present the almost complete, nearly-atomic resolution structure of Carin-1 comprising capsid, portal, and tail machineries at 3.5 Å, 3.8 Å and 3.9 Å, respectively, determined by cryo-electron microscopy (cryo-EM). Our experimental results, combined with AlphaFold2 (AF), allowed us to obtain the nearly-atomic structure of Carin-1 by fitting and refining the AF atomic models in the high resolution cryo-EM map, skipping the bottleneck of manual building and speeding up the structure determination process. Our structural results highlighted the T7-like nature of Carin1, as well as several novel structural features like the presence of short spikes on the capsid, reminiscent those described for Rhodobacter capsulatus gene transfer agent (RcGTA). This is, to our knowledge, the first time such assembly is described for a bacteriophage, shedding light into the common evolution and shared mechanisms between gene transfer agents and phages. This first full structure determined for a marine podophage allowed to propose an infection mechanism different than the one proposed for the archetypal podophage T7. Oceans play a central role in the carbon cycle on Earth and on the climate regulation (half of the planet's CO2 is absorbed by phytoplankton photosynthesis in the oceans and just as much O2 is liberated). The understanding of the biochemical equilibriums of marine biology represents a major goal for our future. By lysing half of the bacterial population every day, marine bacteriophages are key actors of these equilibriums. Despite their importance, these marine phages have, so far, only been studied a little and, in particular, structural insights are currently lacking, even though they are fundamental for the understanding of the molecular mechanisms of their mode of infection. The structures described in our manuscript allow us to propose an infection mechanism that differs from the one proposed for the terrestrial T7 virus, and might also allow us to, in the future, better understand the way bacteriophages shape the global ecosystem. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16688.map.gz emd_16688.map.gz | 92.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16688-v30.xml emd-16688-v30.xml emd-16688.xml emd-16688.xml | 15.7 KB 15.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16688_fsc.xml emd_16688_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_16688.png emd_16688.png | 37.4 KB | ||
| Filedesc metadata |  emd-16688.cif.gz emd-16688.cif.gz | 5.6 KB | ||
| Others |  emd_16688_half_map_1.map.gz emd_16688_half_map_1.map.gz emd_16688_half_map_2.map.gz emd_16688_half_map_2.map.gz | 75.4 MB 75.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16688 http://ftp.pdbj.org/pub/emdb/structures/EMD-16688 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16688 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16688 | HTTPS FTP |
-Validation report
| Summary document |  emd_16688_validation.pdf.gz emd_16688_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16688_full_validation.pdf.gz emd_16688_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_16688_validation.xml.gz emd_16688_validation.xml.gz | 17.4 KB | Display | |
| Data in CIF |  emd_16688_validation.cif.gz emd_16688_validation.cif.gz | 22.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16688 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16688 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16688 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16688 | HTTPS FTP |
-Related structure data
| Related structure data |  8ck0MC  8cjzC  8ck1C M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16688.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16688.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.35 Å | ||||||||||||||||||||||||||||||||||||
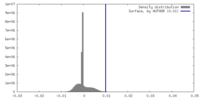
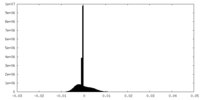
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_16688_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
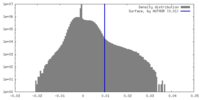
| Density Histograms |
-Half map: #1
| File | emd_16688_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
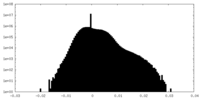
| Density Histograms |
- Sample components
Sample components
-Entire : Bacteriophage sp. Cobetia Marina Virus Carin1
| Entire | Name: Bacteriophage sp. Cobetia Marina Virus Carin1 |
|---|---|
| Components |
|
-Supramolecule #1: Bacteriophage sp.
| Supramolecule | Name: Bacteriophage sp. / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / Details: Cobetia Marina Virus Carin1 / NCBI-ID: 3801 / Sci species name: Bacteriophage sp. / Virus type: VIRION / Virus isolate: OTHER / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Cobetia marina (bacteria) / Strain: DSM 4741 Cobetia marina (bacteria) / Strain: DSM 4741 |
| Virus shell | Shell ID: 1 / Name: Icosahedral capsid / Diameter: 700.0 Å / T number (triangulation number): 7 |
-Macromolecule #1: Portal protein
| Macromolecule | Name: Portal protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Bacteriophage sp. (virus) Bacteriophage sp. (virus) |
| Molecular weight | Theoretical: 62.227227 KDa |
| Sequence | String: MKVETKRQRY NKRLHALKTE RSSWEGSWRD ISDHIQPWRS RFCMSDSNKG NRHNKALVDD TPFKASRTLA SGMMSGLTSP ARPWFKLAI DNPELMESAA VKQWLYLVTQ RMQSVMSKSN LYNQLHQLYG ELGTFGTSCI LIVEDKEDVI RAIPMTVGTY Y LATSSRGM ...String: MKVETKRQRY NKRLHALKTE RSSWEGSWRD ISDHIQPWRS RFCMSDSNKG NRHNKALVDD TPFKASRTLA SGMMSGLTSP ARPWFKLAI DNPELMESAA VKQWLYLVTQ RMQSVMSKSN LYNQLHQLYG ELGTFGTSCI LIVEDKEDVI RAIPMTVGTY Y LATSSRGM VDTIYREMRK TVRQLVQEFG LKNCSNSVQT MYDNDNLEAW VEVVHLIEPN DERNEGKLDS KNKPYRSVYY ES GCTEDKL LRESGFDEFP GVAPRWEVLG DDVYGYGPGH AALGPSRSLQ VMQKKKAQAV EKQINPPMNV PSSSKSKPFS LLP GSLNYY DAAMGGQQMA QPSQSVNLNL NSIMEDIVAH QTSIKETFYA DLFMMIANDQ RSNITAREIQ ERHEEKLLAL GPVV ERLDT EALDPMIDRV FGIMMRGGHL PPPPEEIQGV DLNVQYISVM AQAQKLVGIG AIERITSFVG NLAGADPSAL DKLNI DQAI DQYSNSVSAP PDIIRTDDEV AEIRQARAQQ QQQAQAMEMA QQAAQGAKLL SETDAGTGQN GLSAMMQQMG LA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Component - Concentration: 1.0 x / Component - Name: PBS |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER/RHODIUM / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Details: 25mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 36000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)