[English] 日本語
 Yorodumi
Yorodumi- EMDB-15973: Cryo-EM structure of the Photosystem II - LHCII supercomplex from... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Photosystem II - LHCII supercomplex from Chlorella ohadi | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Green alga / PSII / C.ohadi / membrane protein / Cryo-EM / PHOTOSYNTHESIS | |||||||||
| Function / homology |  Function and homology information Function and homology informationdiacylglycerol kinase (ATP) / ATP-dependent diacylglycerol kinase activity / photosynthesis, light harvesting / photosystem II oxygen evolving complex / photosystem II assembly / oxygen evolving activity / photosystem II stabilization / photosystem II / photosystem II reaction center / : ...diacylglycerol kinase (ATP) / ATP-dependent diacylglycerol kinase activity / photosynthesis, light harvesting / photosystem II oxygen evolving complex / photosystem II assembly / oxygen evolving activity / photosystem II stabilization / photosystem II / photosystem II reaction center / : / photosystem I / oxidoreductase activity, acting on diphenols and related substances as donors, oxygen as acceptor / photosynthetic electron transport chain / response to herbicide / photosystem II / plastid / extrinsic component of membrane / chlorophyll binding / photosynthesis, light reaction / electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity / phosphate ion binding / chloroplast thylakoid membrane / photosynthetic electron transport in photosystem II / photosynthesis / chloroplast / electron transfer activity / protein stabilization / iron ion binding / calcium ion binding / heme binding / RNA binding / ATP binding / membrane / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Chlorella ohadii (plant) Chlorella ohadii (plant) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.73 Å | |||||||||
 Authors Authors | Fadeeva M / Klaiman D / Caspy I / Nelson N | |||||||||
| Funding support |  Israel, 1 items Israel, 1 items
| |||||||||
 Citation Citation |  Journal: Cells / Year: 2023 Journal: Cells / Year: 2023Title: Structure of Photosystem II Reveals Protective Mechanisms against Environmental Stress. Authors: Maria Fadeeva / Daniel Klaiman / Ido Caspy / Nathan Nelson /  Abstract: Green alga is known for its ability to carry out photosynthesis under harsh conditions. Using cryogenic electron microscopy (cryoEM), we obtained a high-resolution structure of PSII at 2.72 Å. This ...Green alga is known for its ability to carry out photosynthesis under harsh conditions. Using cryogenic electron microscopy (cryoEM), we obtained a high-resolution structure of PSII at 2.72 Å. This structure revealed 64 subunits, which encompassed 386 chlorophylls, 86 carotenoids, four plastoquinones, and several structural lipids. At the luminal side of PSII, a unique subunit arrangement was observed to protect the oxygen-evolving complex. This arrangement involved PsbO (OEE1), PsbP (OEE2), PsbB, and PsbU (a homolog of plant OEE3). PsbU interacted with PsbO, PsbC, and PsbP, thereby stabilizing the shield of the oxygen-evolving complex. Significant changes were also observed at the stromal electron acceptor side. PsbY, identified as a transmembrane helix, was situated alongside PsbF and PsbE, which enclosed cytochrome . Supported by the adjacent C-terminal helix of Psb10, these four transmembrane helices formed a bundle that shielded cytochrome from the surrounding solvent. Moreover, the bulk of Psb10 formed a protective cap, which safeguarded the quinone site and likely contributed to the stacking of PSII complexes. Based on our findings, we propose a protective mechanism that prevents Q (plastoquinone B) from becoming fully reduced. This mechanism offers insights into the regulation of electron transfer within PSII. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15973.map.gz emd_15973.map.gz | 443.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15973-v30.xml emd-15973-v30.xml emd-15973.xml emd-15973.xml | 49.8 KB 49.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15973_fsc.xml emd_15973_fsc.xml | 17.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_15973.png emd_15973.png | 33.2 KB | ||
| Others |  emd_15973_half_map_1.map.gz emd_15973_half_map_1.map.gz emd_15973_half_map_2.map.gz emd_15973_half_map_2.map.gz | 382.7 MB 382.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15973 http://ftp.pdbj.org/pub/emdb/structures/EMD-15973 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15973 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15973 | HTTPS FTP |
-Validation report
| Summary document |  emd_15973_validation.pdf.gz emd_15973_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15973_full_validation.pdf.gz emd_15973_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_15973_validation.xml.gz emd_15973_validation.xml.gz | 25.4 KB | Display | |
| Data in CIF |  emd_15973_validation.cif.gz emd_15973_validation.cif.gz | 33.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15973 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15973 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15973 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15973 | HTTPS FTP |
-Related structure data
| Related structure data |  8bd3MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15973.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15973.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.9216 Å | ||||||||||||||||||||||||||||||||||||
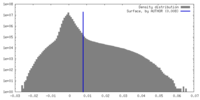
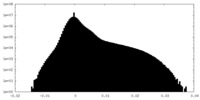
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_15973_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
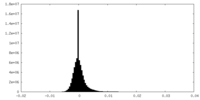
| Density Histograms |
-Half map: #1
| File | emd_15973_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
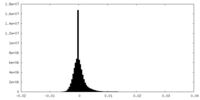
| Density Histograms |
- Sample components
Sample components
+Entire : Chlorella ohadii Photosystem II
+Supramolecule #1: Chlorella ohadii Photosystem II
+Macromolecule #1: Chlorophyll a-b binding protein, chloroplastic
+Macromolecule #2: Chlorophyll a-b binding protein, chloroplastic
+Macromolecule #3: Chlorophyll a-b binding of LHCII
+Macromolecule #4: Chlorophyll a-b binding of LHCII
+Macromolecule #5: Photosystem II protein D1
+Macromolecule #6: Photosystem II CP47 reaction center protein
+Macromolecule #7: Photosystem II CP43 reaction center protein
+Macromolecule #8: Photosystem II D2 protein
+Macromolecule #9: Cytochrome b559 subunit alpha
+Macromolecule #10: Cytochrome b559 subunit beta
+Macromolecule #11: Photosystem II reaction center protein H
+Macromolecule #12: Photosystem II reaction center protein I
+Macromolecule #13: Photosystem II reaction center protein J
+Macromolecule #14: Photosystem II reaction center protein K
+Macromolecule #15: Photosystem II reaction center protein L
+Macromolecule #16: Photosystem II reaction center protein M
+Macromolecule #17: Chloroplast oxygen-evolving enhancer protein 1
+Macromolecule #18: Chlorophyll a b binding CP29
+Macromolecule #19: Chlorophyll a b-binding CP26
+Macromolecule #20: Photosystem II reaction center protein T
+Macromolecule #21: Photosystem II reaction center protein Ycf12
+Macromolecule #22: Photosystem II reaction center W protein
+Macromolecule #23: Photosystem II reaction center protein X
+Macromolecule #24: Multifunctional fusion protein
+Macromolecule #25: Photosystem II reaction center protein Z
+Macromolecule #26: Chloroplast oxygen-evolving enhancer protein 3
+Macromolecule #27: Chloroplast PsbY
+Macromolecule #28: Chloroplast photosystem II 10 kDa protein
+Macromolecule #29: Photosystem II oxygen evolving enhancer 2
+Macromolecule #30: CHLOROPHYLL B
+Macromolecule #31: CHLOROPHYLL A
+Macromolecule #32: (3R,3'R,6S)-4,5-DIDEHYDRO-5,6-DIHYDRO-BETA,BETA-CAROTENE-3,3'-DIOL
+Macromolecule #33: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE
+Macromolecule #34: 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE
+Macromolecule #35: (3R)-beta,beta-caroten-3-ol
+Macromolecule #36: (1R,3R)-6-{(3E,5E,7E,9E,11E,13E,15E,17E)-18-[(1S,4R,6R)-4-HYDROXY...
+Macromolecule #37: (3S,5R,6S,3'S,5'R,6'S)-5,6,5',6'-DIEPOXY-5,6,5',6'- TETRAHYDRO-BE...
+Macromolecule #38: 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL
+Macromolecule #39: CA-MN4-O5 CLUSTER
+Macromolecule #40: FE (II) ION
+Macromolecule #41: CHLORIDE ION
+Macromolecule #42: PHEOPHYTIN A
+Macromolecule #43: BETA-CAROTENE
+Macromolecule #44: 2,3-DIMETHYL-5-(3,7,11,15,19,23,27,31,35-NONAMETHYL-2,6,10,14,18,...
+Macromolecule #45: DIGALACTOSYL DIACYL GLYCEROL (DGDG)
+Macromolecule #46: BICARBONATE ION
+Macromolecule #47: PROTOPORPHYRIN IX CONTAINING FE
+Macromolecule #48: DODECYL-ALPHA-D-MALTOSIDE
+Macromolecule #49: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL |
|---|---|
| Buffer | pH: 6 / Details: 20 mM MES NaOH pH 6 0.1% alfaDM |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 12 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 293 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 21341 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)