[English] 日本語
 Yorodumi
Yorodumi- EMDB-15243: Mycobacterium tuberculosis ClpC1 hexamer structure bound to the n... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mycobacterium tuberculosis ClpC1 hexamer structure bound to the natural product antibiotic ecumicin (class 2) | ||||||||||||
 Map data Map data | Mycobacterium tuberculosis ClpC1 with added Ecumicin, Class 2 | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | hexamer / tuberculosis / drug target / protein quality control / CHAPERONE | ||||||||||||
| Biological species |  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.59 Å | ||||||||||||
 Authors Authors | Felix J / Fraga H / Gragera M / Bueno T / Weinhaeupl K | ||||||||||||
| Funding support | European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2022 Journal: J Biol Chem / Year: 2022Title: Structure of the drug target ClpC1 unfoldase in action provides insights on antibiotic mechanism of action. Authors: Katharina Weinhäupl / Marcos Gragera / M Teresa Bueno-Carrasco / Rocío Arranz / Olga Krandor / Tatos Akopian / Raquel Soares / Eric Rubin / Jan Felix / Hugo Fraga /     Abstract: The unfoldase ClpC1 is one of the most exciting drug targets against tuberculosis. This AAA+ unfoldase works in cooperation with the ClpP1P2 protease and is the target of at least four natural ...The unfoldase ClpC1 is one of the most exciting drug targets against tuberculosis. This AAA+ unfoldase works in cooperation with the ClpP1P2 protease and is the target of at least four natural product antibiotics: cyclomarin, ecumicin, lassomycin, and rufomycin. Although these molecules are promising starting points for drug development, their mechanisms of action remain largely unknown. Taking advantage of a middle domain mutant, we determined the first structure of Mycobacterium tuberculosis ClpC1 in its apo, cyclomarin-, and ecumicin-bound states via cryo-EM. The obtained structure displays features observed in other members of the AAA+ family and provides a map for further drug development. While the apo and cyclomarin-bound structures are indistinguishable and have N-terminal domains that are invisible in their respective EM maps, around half of the ecumicin-bound ClpC1 particles display three of their six N-terminal domains in an extended conformation. Our structural observations suggest a mechanism where ecumicin functions by mimicking substrate binding, leading to ATPase activation and changes in protein degradation profile. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15243.map.gz emd_15243.map.gz | 56.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15243-v30.xml emd-15243-v30.xml emd-15243.xml emd-15243.xml | 16.3 KB 16.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15243_fsc.xml emd_15243_fsc.xml | 8.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_15243.png emd_15243.png | 51.7 KB | ||
| Others |  emd_15243_half_map_1.map.gz emd_15243_half_map_1.map.gz emd_15243_half_map_2.map.gz emd_15243_half_map_2.map.gz | 55.4 MB 55.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15243 http://ftp.pdbj.org/pub/emdb/structures/EMD-15243 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15243 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15243 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15243.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15243.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium tuberculosis ClpC1 with added Ecumicin, Class 2 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.71 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_15243_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
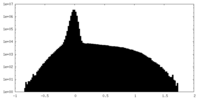
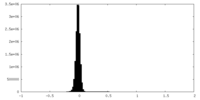
| Density Histograms |
-Half map: #2
| File | emd_15243_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
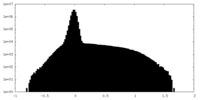
| Density Histograms |
- Sample components
Sample components
-Entire : Mycobacterium tuberculosis ClpC1 bound to the natural product ant...
| Entire | Name: Mycobacterium tuberculosis ClpC1 bound to the natural product antibiotic Ecumicin |
|---|---|
| Components |
|
-Supramolecule #1: Mycobacterium tuberculosis ClpC1 bound to the natural product ant...
| Supramolecule | Name: Mycobacterium tuberculosis ClpC1 bound to the natural product antibiotic Ecumicin type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 567 KDa |
-Macromolecule #1: Mycobacterium tuberculosis ClpC1 with added Ecumicin, Class 2
| Macromolecule | Name: Mycobacterium tuberculosis ClpC1 with added Ecumicin, Class 2 type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MFERFTDRAR RVVVLAQEEA RMLNHNYIGT EHILLGLIHE GEGVAAKSLE SLGISLEGVR SQVEEIIGQ GQQAPSGHIP FTPRAKKVLE LSLREALQLG HNYIGTEHIL LGLIREGEGV A AQVLVKLG AELTRVRQQV IQLLSGYQGK EAAEAGTGGR GGESGSPSTS ...String: MFERFTDRAR RVVVLAQEEA RMLNHNYIGT EHILLGLIHE GEGVAAKSLE SLGISLEGVR SQVEEIIGQ GQQAPSGHIP FTPRAKKVLE LSLREALQLG HNYIGTEHIL LGLIREGEGV A AQVLVKLG AELTRVRQQV IQLLSGYQGK EAAEAGTGGR GGESGSPSTS LVLDQFGRNL TA AAMEGKL DPVIGREKEI ERVMQVLSRR TKNNPVLIGE PGVGKTAVVE GLAQAIVHGE VPE TLKDKQ LYTLDLGSLV AGSRYRGDFE ERLKKVLKEI NTRGDIILFI DALHTLVGAG AAEG AIDAA SILKPKLARG ELQTIGATTL DEYRKYIEKD AALERRFQPV QVGEPTVEHT IEILK GLRD RYEAHHRVSI TDAAMVAAAT LADRYINDRF LPDKAIDLID EAGARMRIRR MTAPPD LRE FDEKIAEARR EKESAIDAQD FEKAASLRDR EKTLVAQRAE REKQWRSGDL DVVAEVD DE QIAEVLGNWT GIPVFKLTEA ETTRLLRMEE ELHKRIIGQE DAVKAVSKAI RRTRAGLK D PKRPSGSFIF AGPSGVGKTE LSKALANFLF GDDDALIQID MGEFHDRFTA SRLFGAPPG YVGYEEGGQL TEKVRRKPFS VVLFDAIEKA HQEIYNSLLQ VLEDGRLTDG QGRTVDFKNT VLIFTSNLG TSDISKPVGL GFSKGGGEND YERMKQKVND ELKKHFRPEF LNRIDDIIVF H QLTREEII RMVDLMISRV AGQLKSKDMA LVLTDAAKAL LAKRGFDPVL GARPLRRTIQ RE IEDQLSE KILFEEVGPG QVVTVDVDNW DGEGPGEDAV FTFTGTRKPP AEPDLAKAGA HSA GGPEPA ARLEHHHHHH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.4 Details: Hepes pH 7.4 50 mM, NaCl 100 mM, 10 mM MgCl2, ATP 1mM |
| Vitrification | Cryogen name: ETHANE |
| Details | 10 microM ClpC1 with 30 microM Ecumicin |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average exposure time: 40.0 sec. / Average electron dose: 32.2 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: OTHER / Nominal defocus max: 3.1 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X