[English] 日本語
 Yorodumi
Yorodumi- EMDB-13906: Cryo-EM structure of Tn4430 TnpA transposase from Tn3 family in c... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Tn4430 TnpA transposase from Tn3 family in complex with 100 bp long transposon end DNA | |||||||||||||||
 Map data Map data | composite map created in phenix_combine_phocused_map from consensus refinement of dimer and aligned body 2. The map was used for model refinement. | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | DNA transposition / Tn3 family / antibiotic resistance / protein metamorphosis / RECOMBINATION | |||||||||||||||
| Function / homology | Tn3 transposase DDE domain / Domain of unknown function DUF4158 / : / Tn3 transposase DDE domain / Domain of unknown function (DUF4158) / transposase activity / DNA transposition / DNA binding / Transposase for transposon Tn4430 Function and homology information Function and homology information | |||||||||||||||
| Biological species |  | |||||||||||||||
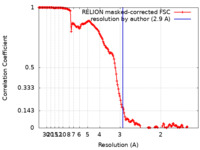
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||||||||
 Authors Authors | Shkumatov AV / Oger CA / Aryanpour N / Hallet BF / Efremov RG | |||||||||||||||
| Funding support |  Belgium, 4 items Belgium, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural insight into Tn3 family transposition mechanism. Authors: Alexander V Shkumatov / Nicolas Aryanpour / Cédric A Oger / Gérôme Goossens / Bernard F Hallet / Rouslan G Efremov /  Abstract: Transposons are diverse mobile genetic elements that play the critical role as genome architects in all domains of life. Tn3 is a widespread family and among the first identified bacterial ...Transposons are diverse mobile genetic elements that play the critical role as genome architects in all domains of life. Tn3 is a widespread family and among the first identified bacterial transposons famed for their contribution to the dissemination of antibiotic resistance. Transposition within this family is mediated by a large TnpA transposase, which facilitates both transposition and target immunity. Howtever, a structural framework required for understanding the mechanism of TnpA transposition is lacking. Here, we describe the cryo-EM structures of TnpA from Tn4430 in the apo form and paired with transposon ends before and after DNA cleavage and strand transfer. We show that TnpA has an unusual architecture and exhibits a family specific regulatory mechanism involving metamorphic refolding of the RNase H-like catalytic domain. The TnpA structure, constrained by a double dimerization interface, creates a peculiar topology that suggests a specific role for the target DNA in transpososome assembly and activation. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13906.map.gz emd_13906.map.gz | 191.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13906-v30.xml emd-13906-v30.xml emd-13906.xml emd-13906.xml | 24.5 KB 24.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13906_fsc.xml emd_13906_fsc.xml | 15.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_13906.png emd_13906.png | 102.2 KB | ||
| Masks |  emd_13906_msk_1.map emd_13906_msk_1.map | 325 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-13906.cif.gz emd-13906.cif.gz | 7.3 KB | ||
| Others |  emd_13906_additional_1.map.gz emd_13906_additional_1.map.gz emd_13906_half_map_1.map.gz emd_13906_half_map_1.map.gz emd_13906_half_map_2.map.gz emd_13906_half_map_2.map.gz | 6.4 MB 259.6 MB 259.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13906 http://ftp.pdbj.org/pub/emdb/structures/EMD-13906 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13906 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13906 | HTTPS FTP |
-Validation report
| Summary document |  emd_13906_validation.pdf.gz emd_13906_validation.pdf.gz | 842.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_13906_full_validation.pdf.gz emd_13906_full_validation.pdf.gz | 842 KB | Display | |
| Data in XML |  emd_13906_validation.xml.gz emd_13906_validation.xml.gz | 23.4 KB | Display | |
| Data in CIF |  emd_13906_validation.cif.gz emd_13906_validation.cif.gz | 30.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13906 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13906 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13906 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13906 | HTTPS FTP |
-Related structure data
| Related structure data |  7qd4MC  7qd5C  7qd6C  7qd8C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13906.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13906.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | composite map created in phenix_combine_phocused_map from consensus refinement of dimer and aligned body 2. The map was used for model refinement. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.766 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_13906_msk_1.map emd_13906_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
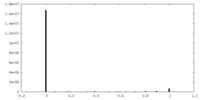
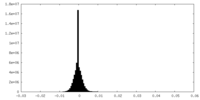
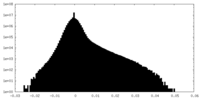
| Density Histograms |
-Additional map: post processed map of body2 used for model...
| File | emd_13906_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | post processed map of body2 used for model building. Not aligned to the map of the dimer | ||||||||||||
| Projections & Slices |
| ||||||||||||
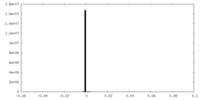
| Density Histograms |
-Half map: half map 1
| File | emd_13906_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_13906_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : TnpA-IR100 complex
| Entire | Name: TnpA-IR100 complex |
|---|---|
| Components |
|
-Supramolecule #1: TnpA-IR100 complex
| Supramolecule | Name: TnpA-IR100 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: hyperactive TnpA mutant S911R in complex with 100bp DNA containing trasnposone recognition sequence. |
|---|
-Supramolecule #2: hyperactive TnpA mutant S911R
| Supramolecule | Name: hyperactive TnpA mutant S911R / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 / Details: hyperactive TnpA mutant S911R |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: DNA substrate
| Supramolecule | Name: DNA substrate / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 / Details: DNA substrate |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Transposase for transposon Tn4430
| Macromolecule | Name: Transposase for transposon Tn4430 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 116.992117 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGVKQLLSEA QRNELMDLSR LTEWDLVTFH TFSKHDLHLI LKHRRGYNRL GFALQLVLIR YPGWSLTEYK DIPQYVVAYV ASQLQIPPE EFLVYAKRGN TLWEHLGEIR TEYGYQNFSS EYKETLLQFL VQQAMDNNNT LYLIEITIST LRKMKVILPA M YVIEDIVW ...String: MGVKQLLSEA QRNELMDLSR LTEWDLVTFH TFSKHDLHLI LKHRRGYNRL GFALQLVLIR YPGWSLTEYK DIPQYVVAYV ASQLQIPPE EFLVYAKRGN TLWEHLGEIR TEYGYQNFSS EYKETLLQFL VQQAMDNNNT LYLIEITIST LRKMKVILPA M YVIEDIVW EAKQQADQKV YSILHDGLVQ EQKDQLDALL LPTINGKSPL AWLKDVPAQP SPESFLKVID RLQFVQKIGL TI DTTKINT NRLRQLARLG SKYEPYAFRR FNEVKRYSML VSFLLEITQD LIDYAIEIHD RLMMNLQTKG KKEQDEIQQA NGK KLNEKI LQFITVCGTL IEAKETGKDA FAALDEVMSW NEMVESVEEA KQLSRPLNYD YLDLLNTRYS YVRRYAPTLL RSLH FRATK SGEPVLQALD TIHELNETGK RKVPHGAPLH FVSNRWQKHV YDDDGNINRH YYELAALTEL RNHIRSGDIF VSGSR HHKA FDDYLIPYDE WNEVSNIPNG LTAPLKAEDY ITDRINRLNE HLEWLSKNSE KLEGVDISQG KLHVERLDRG TPEEAK AFS KLLHSMLPRI KLTDLLIEVA SWTGFHDQFI HASTNQSPDQ EEQNIVLATL MAMGTNIGLT KMAEATPGIS YRQMANA SQ WRMYDDAMVR AQSILVNFQK EQKLSSYWGD GTTSSSDGMR LSIAVRSLHA DSNPHYGTGK GGTIYRFVSD QLSAYHVK V ITTNARDALH VLDGLLHHET DLKIEEHYTD TAGYTDQVFA LTHLLGFRFA PRIRDLADTK LFSIPGGEEY ENVQALLKG KINVKLIKEN YEDIRRLAYS VQTGKVSSAL IMGKLGSYAR QNKLATALGE MGRIEKTLFT LDYISNKAVR RRVQKGLNKG EAINALARI IFFGQRGEFR ERALQDQLQR ARALNIIINA ISVWNTVYME KAVEELKARG EFREDLMPYA WPLGWEHINF L GEYKFEGL HDTGQMNLRP LRIKEPFYSP IRSFLEQKLI SEEDLNSAVD HHHHHH UniProtKB: Transposase for transposon Tn4430 |
-Macromolecule #2: IR100 DNA substrate, none transferred strand
| Macromolecule | Name: IR100 DNA substrate, none transferred strand / type: dna / ID: 2 Details: 100 base pairs internal DNA repeat containing recognition sequence for Tn4430 transposone. Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 30.848781 KDa |
| Sequence | String: (DC)(DG)(DG)(DG)(DG)(DA)(DT)(DC)(DC)(DT) (DC)(DT)(DA)(DG)(DC)(DT)(DC)(DG)(DA)(DG) (DA)(DT)(DG)(DC)(DA)(DT)(DC)(DC)(DA) (DT)(DG)(DG)(DG)(DG)(DG)(DT)(DA)(DC)(DC) (DG) (DC)(DC)(DA)(DG)(DC)(DA) ...String: (DC)(DG)(DG)(DG)(DG)(DA)(DT)(DC)(DC)(DT) (DC)(DT)(DA)(DG)(DC)(DT)(DC)(DG)(DA)(DG) (DA)(DT)(DG)(DC)(DA)(DT)(DC)(DC)(DA) (DT)(DG)(DG)(DG)(DG)(DG)(DT)(DA)(DC)(DC) (DG) (DC)(DC)(DA)(DG)(DC)(DA)(DT)(DT) (DT)(DC)(DG)(DG)(DA)(DA)(DA)(DA)(DA)(DA) (DA)(DC) (DC)(DA)(DC)(DG)(DC)(DT)(DA) (DA)(DG)(DA)(DT)(DC)(DC)(DT)(DC)(DT)(DA) (DG)(DG)(DC) (DT)(DA)(DG)(DC)(DG)(DC) (DG)(DC)(DA)(DA)(DA)(DA)(DT)(DA)(DA)(DT) (DA)(DC)(DA)(DA) |
-Macromolecule #3: IR100 DNA substrate, transferred strand
| Macromolecule | Name: IR100 DNA substrate, transferred strand / type: dna / ID: 3 Details: 100 base pairs internal DNA repeat containing recognition sequence for Tn4430 transposone. Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 30.851658 KDa |
| Sequence | String: (DT)(DT)(DG)(DT)(DA)(DT)(DT)(DA)(DT)(DT) (DT)(DT)(DG)(DC)(DG)(DC)(DG)(DC)(DT)(DA) (DG)(DC)(DC)(DT)(DA)(DG)(DA)(DG)(DG) (DA)(DT)(DC)(DT)(DT)(DA)(DG)(DC)(DG)(DT) (DG) (DG)(DT)(DT)(DT)(DT)(DT) ...String: (DT)(DT)(DG)(DT)(DA)(DT)(DT)(DA)(DT)(DT) (DT)(DT)(DG)(DC)(DG)(DC)(DG)(DC)(DT)(DA) (DG)(DC)(DC)(DT)(DA)(DG)(DA)(DG)(DG) (DA)(DT)(DC)(DT)(DT)(DA)(DG)(DC)(DG)(DT) (DG) (DG)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DC)(DC)(DG)(DA)(DA)(DA)(DT)(DG)(DC)(DT) (DG)(DG) (DC)(DG)(DG)(DT)(DA)(DC)(DC) (DC)(DC)(DC)(DA)(DT)(DG)(DG)(DA)(DT)(DG) (DC)(DA)(DT) (DC)(DT)(DC)(DG)(DA)(DG) (DC)(DT)(DA)(DG)(DA)(DG)(DG)(DA)(DT)(DC) (DC)(DC)(DC)(DG) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.15 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 50 mM HEPES (pH 7.5), 100 mM NaCl, 30 mM L-Arg HCL |
| Grid | Model: Quantifoil R2/1 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-40 / Number grids imaged: 1 / Number real images: 4756 / Average exposure time: 3.0 sec. / Average electron dose: 62.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 6.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: JEOL CRYOSPECPORTER / Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 28.8 |
|---|---|
| Output model |  PDB-7qd4: |
 Movie
Movie Controller
Controller






 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)