+Search query
-Structure paper
| Title | Lipids modulate the open probability of RyR1 under cryo-EM conditions. |
|---|---|
| Journal, issue, pages | Structure, Vol. 33, Issue 12, Page 2029-22040.e3, Year 2025 |
| Publish date | Dec 4, 2025 |
 Authors Authors | Chenyao Li / Rouslan G Efremov /  |
| PubMed Abstract | Ryanodine receptors (RyRs) are intracellular tetrameric ion channels responsible for Ca release from the sarcoplasmic and endoplasmic reticulum. Ryanodine receptor 1 (RyR1) isoform, critical for ...Ryanodine receptors (RyRs) are intracellular tetrameric ion channels responsible for Ca release from the sarcoplasmic and endoplasmic reticulum. Ryanodine receptor 1 (RyR1) isoform, critical for muscle contraction, has been studied most extensively. While cryoelectron microscopy (cryo-EM) has been instrumental in revealing near-atomic details of RyR gating mechanisms, the open probability of RyR1 under cryo-EM conditions is notably lower than that observed in electrophysiological studies. Here, we present a cryo-EM study examining the open probability of RyR1 solubilized in CHAPS with varying lipid concentrations. We found that increasing lipid concentration from 0.001% to 0.05% raised the RyR1 open probability from 16% to 84%, whereas RyR1 reconstituted into lipid nanodiscs remained closed. We modeled 72 lipid molecules in the map reconstructed at the highest lipid concentration. These findings demonstrate the important role of lipids in modulating the open fraction of solubilized RyR1 channels under cryo-EM conditions and suggest optimal lipid mimetics for structural studies of RyR1 gating. |
 External links External links |  Structure / Structure /  PubMed:41027431 PubMed:41027431 |
| Methods | EM (single particle) |
| Resolution | 3.3 - 3.6 Å |
| Structure data | EMDB-52085, PDB-9heo: EMDB-52091, PDB-9heq: EMDB-53834, PDB-9r8o: EMDB-53924, PDB-9rcw: |
| Chemicals |  ChemComp-ZN:  ChemComp-ATP: 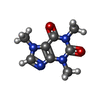 ChemComp-CFF:  ChemComp-CA:  ChemComp-POV: |
| Source |
|
 Keywords Keywords | TRANSPORT PROTEIN / Ion channel / Ca2+ / tetramer |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers












