+Search query
-Structure paper
| Title | Molecular basis of SLC19A1-mediated folate and cyclic dinucleotide transport. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 16, Issue 1, Page 3146, Year 2025 |
| Publish date | Apr 2, 2025 |
 Authors Authors | Qixiang Zhang / Xuyuan Zhang / Kexin Liu / Yalan Zhu / Xiaohua Nie / Junxiao Ma / Panpan Sun / Zhaolong Li / Yina Gao / Songqing Liu / Ang Gao / Liguo Zhang / Pu Gao /  |
| PubMed Abstract | The solute carrier protein SLC19A1 is crucial for transporting folate nutrients, antifolate chemotherapeutics, and more recently cyclic dinucleotides (CDNs) immune transmitters, influencing various ...The solute carrier protein SLC19A1 is crucial for transporting folate nutrients, antifolate chemotherapeutics, and more recently cyclic dinucleotides (CDNs) immune transmitters, influencing various physiological and pathological processes. While the inward-open state of human SLC19A1 (hSLC19A1) has been previously described, key aspects regarding its conformational dynamics, substrate selectivity, and precise mechanisms underlying CDNs transport remain elusive. Using an antibody-facilitated conformation screening strategy, we present cryo-electron microscopy structures of hSLC19A1 in its outward-open state with and without bound substrates, revealing detailed mechanisms of substrate recognition and conformational changes during transport. We identify both general and specific features for folate/antifolate recognition, including an SLC19A1-specific pocket for accommodating γ-carboxylate-modified antifolates. Intriguingly, CDNs bind as monomers within the canonical pocket of outward-open hSLC19A1, contrasting with dimeric binding in inward-open structures. Together with functional assays, these findings provide a framework for developing antifolate drugs and CDN-targeted therapies, advancing our understanding of SLC19A1's physiological and therapeutic functions. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:40175380 / PubMed:40175380 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.94 - 3.74 Å |
| Structure data | EMDB-61690, PDB-9joz: EMDB-61756, PDB-9jri: EMDB-61758, PDB-9jrk: EMDB-61759, PDB-9jrl: EMDB-61760, PDB-9jrm: EMDB-61761, PDB-9jrn: |
| Chemicals | 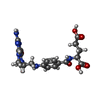 ChemComp-C2F: 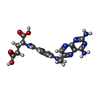 ChemComp-MTX: 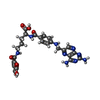 ChemComp-COP: 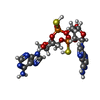 ChemComp-GJF:  ChemComp-TPP: |
| Source |
|
 Keywords Keywords | MEMBRANE PROTEIN / outward-open hSLC19A1 |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers















 homo sapiens (human)
homo sapiens (human)
