+Search query
-Structure paper
| Title | A mini-hairpin shaped nascent peptide blocks translation termination by a distinct mechanism. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 16, Issue 1, Page 2323, Year 2025 |
| Publish date | Mar 8, 2025 |
 Authors Authors | Yushin Ando / Akinao Kobo / Tatsuya Niwa / Ayako Yamakawa / Suzuna Konoma / Yuki Kobayashi / Osamu Nureki / Hideki Taguchi / Yuzuru Itoh / Yuhei Chadani /  |
| PubMed Abstract | Protein synthesis by ribosomes produces functional proteins but also serves diverse regulatory functions, which depend on the coding amino acid sequences. Certain nascent peptides interact with the ...Protein synthesis by ribosomes produces functional proteins but also serves diverse regulatory functions, which depend on the coding amino acid sequences. Certain nascent peptides interact with the ribosome exit tunnel to arrest translation and modulate themselves or the expression of downstream genes. However, a comprehensive understanding of the mechanisms of such ribosome stalling and its regulation remains elusive. In this study, we systematically screen for unidentified ribosome arrest peptides through phenotypic evaluation, proteomics, and mass spectrometry analyses, leading to the discovery of the arrest peptides PepNL and NanCL in E. coli. Our cryo-EM study on PepNL reveals a distinct arrest mechanism, in which the N-terminus of PepNL folds back towards the tunnel entrance to prevent the catalytic GGQ motif of the release factor from accessing the peptidyl transferase center, causing translation arrest at the UGA stop codon. Furthermore, unlike sensory arrest peptides that require an arrest inducer, PepNL uses tryptophan as an arrest inhibitor, where Trp-tRNA reads through the stop codon. Our findings illuminate the mechanism and regulatory framework of nascent peptide-induced translation arrest, paving the way for exploring regulatory nascent peptides. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:40057501 / PubMed:40057501 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.9 Å |
| Structure data | EMDB-60473, PDB-8ztu: EMDB-60474, PDB-8ztv: |
| Chemicals |  ChemComp-MG:  ChemComp-K:  ChemComp-SPD: 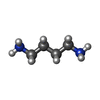 ChemComp-PUT:  ChemComp-ZN: |
| Source |
|
 Keywords Keywords | RIBOSOME / Translation arrest / Nascent peptide / Release factor / tRNA |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers








