+Search query
-Structure paper
| Title | The structural basis of odorant recognition in insect olfactory receptors. |
|---|---|
| Journal, issue, pages | Nature, Vol. 597, Issue 7874, Page 126-131, Year 2021 |
| Publish date | Aug 4, 2021 |
 Authors Authors | Josefina Del Mármol / Mackenzie A Yedlin / Vanessa Ruta /  |
| PubMed Abstract | Olfactory systems must detect and discriminate amongst an enormous variety of odorants. To contend with this challenge, diverse species have converged on a common strategy in which odorant identity ...Olfactory systems must detect and discriminate amongst an enormous variety of odorants. To contend with this challenge, diverse species have converged on a common strategy in which odorant identity is encoded through the combinatorial activation of large families of olfactory receptors, thus allowing a finite number of receptors to detect a vast chemical world. Here we offer structural and mechanistic insight into how an individual olfactory receptor can flexibly recognize diverse odorants. We show that the olfactory receptor MhOR5 from the jumping bristletail Machilis hrabei assembles as a homotetrameric odorant-gated ion channel with broad chemical tuning. Using cryo-electron microscopy, we elucidated the structure of MhOR5 in multiple gating states, alone and in complex with two of its agonists-the odorant eugenol and the insect repellent DEET. Both ligands are recognized through distributed hydrophobic interactions within the same geometrically simple binding pocket located in the transmembrane region of each subunit, suggesting a structural logic for the promiscuous chemical sensitivity of this receptor. Mutation of individual residues lining the binding pocket predictably altered the sensitivity of MhOR5 to eugenol and DEET and broadly reconfigured the receptor's tuning. Together, our data support a model in which diverse odorants share the same structural determinants for binding, shedding light on the molecular recognition mechanisms that ultimately endow the olfactory system with its immense discriminatory capacity. |
 External links External links |  Nature / Nature /  PubMed:34349260 / PubMed:34349260 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.9 - 3.3 Å |
| Structure data | EMDB-23372: The structure of the insect olfactory receptor OR5 from Machilis hrabei. EMDB-23374: The structure of the insect olfactory receptor OR5 from Machilis hrabei in complex with eugenol. EMDB-23375: The structure of the insect olfactory receptor OR5 from Machilis hrabei in complex with DEET. |
| Chemicals | 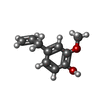 ChemComp-EOL: 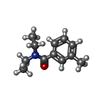 ChemComp-DE3: |
| Source |
|
 Keywords Keywords | MEMBRANE PROTEIN / olfactory receptor / ion channel / eugenol / DEET |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers









 machilis hrabei (insect)
machilis hrabei (insect)