+Search query
-Structure paper
| Title | Structural basis for a reciprocal regulation between SCF and CSN. |
|---|---|
| Journal, issue, pages | Cell Rep, Vol. 2, Issue 3, Page 616-627, Year 2012 |
| Publish date | Sep 27, 2012 |
 Authors Authors | Radoslav I Enchev / Daniel C Scott / Paula C A da Fonseca / Anne Schreiber / Julie K Monda / Brenda A Schulman / Matthias Peter / Edward P Morris /  |
| PubMed Abstract | Skp1-Cul1-Fbox (SCF) E3 ligases are activated by ligation to the ubiquitin-like protein Nedd8, which is reversed by the deneddylating Cop9 signalosome (CSN). However, CSN also promotes SCF substrate ...Skp1-Cul1-Fbox (SCF) E3 ligases are activated by ligation to the ubiquitin-like protein Nedd8, which is reversed by the deneddylating Cop9 signalosome (CSN). However, CSN also promotes SCF substrate turnover through unknown mechanisms. Through biochemical and electron microscopy analyses, we determined molecular models of CSN complexes with SCF(Skp2/Cks1) and SCF(Fbw7) and found that CSN occludes both SCF functional sites-the catalytic Rbx1-Cul1 C-terminal domain and the substrate receptor. Indeed, CSN binding prevents SCF interactions with E2 enzymes and a ubiquitination substrate, and it inhibits SCF-catalyzed ubiquitin chain formation independent of deneddylation. Importantly, CSN prevents neddylation of the bound cullin, unless binding of a ubiquitination substrate triggers SCF dissociation and neddylation. Taken together, the results provide a model for how reciprocal regulation sensitizes CSN to the SCF assembly state and inhibits a catalytically competent SCF until a ubiquitination substrate drives its own degradation by displacing CSN, thereby promoting cullin neddylation and substrate ubiquitination. |
 External links External links |  Cell Rep / Cell Rep /  PubMed:22959436 / PubMed:22959436 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 25.0 Å |
| Structure data | 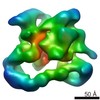 EMDB-2173: 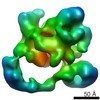 EMDB-2174: 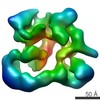 EMDB-2175:  EMDB-2176:  EMDB-2177: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers



 Homo sapiens (human)
Homo sapiens (human)