+Search query
-Structure paper
| Title | Ligand recognition and biased agonism of the D1 dopamine receptor. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 13, Issue 1, Page 3186, Year 2022 |
| Publish date | Jun 8, 2022 |
 Authors Authors | Xiao Teng / Sijia Chen / Yingying Nie / Peng Xiao / Xiao Yu / Zhenhua Shao / Sanduo Zheng /  |
| PubMed Abstract | Dopamine receptors are widely distributed in the central nervous system and are important therapeutic targets for treatment of various psychiatric and neurological diseases. Here, we report three ...Dopamine receptors are widely distributed in the central nervous system and are important therapeutic targets for treatment of various psychiatric and neurological diseases. Here, we report three cryo-electron microscopy structures of the D1 dopamine receptor (D1R)-Gs complex bound to two agonists, fenoldopam and tavapadon, and a positive allosteric modulator LY3154207. The structure reveals unusual binding of two fenoldopam molecules, one to the orthosteric binding pocket (OBP) and the other to the extended binding pocket (EBP). In contrast, one elongated tavapadon molecule binds to D1R, extending from OBP to EBP. Moreover, LY3154207 stabilizes the second intracellular loop of D1R in an alpha helical conformation to efficiently engage the G protein. Through a combination of biochemical, biophysical and cellular assays, we further show that the broad conformation stabilized by two fenoldopam molecules and interaction between TM5 and the agonist are important for biased signaling of D1R. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:35676276 / PubMed:35676276 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.0 - 3.3 Å |
| Structure data | EMDB-32964, PDB-7x2c: EMDB-32965, PDB-7x2d: EMDB-32966, PDB-7x2f: |
| Chemicals |  ChemComp-CLR:  ChemComp-G3C: 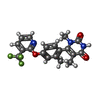 ChemComp-86W: 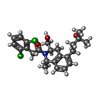 ChemComp-G4C:  ChemComp-LDP: |
| Source |
|
 Keywords Keywords | SIGNALING PROTEIN / GPCR / dopamine receptor / mini-Gs / membrane protein / tavapadon |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers








 homo sapiens (human)
homo sapiens (human)