+Search query
-Structure paper
| Title | The Ccr4-Not complex monitors the translating ribosome for codon optimality. |
|---|---|
| Journal, issue, pages | Science, Vol. 368, Issue 6488, Year 2020 |
| Publish date | Apr 17, 2020 |
 Authors Authors | Robert Buschauer / Yoshitaka Matsuo / Takato Sugiyama / Ying-Hsin Chen / Najwa Alhusaini / Thomas Sweet / Ken Ikeuchi / Jingdong Cheng / Yasuko Matsuki / Risa Nobuta / Andrea Gilmozzi / Otto Berninghausen / Petr Tesina / Thomas Becker / Jeff Coller / Toshifumi Inada / Roland Beckmann /    |
| PubMed Abstract | Control of messenger RNA (mRNA) decay rate is intimately connected to translation elongation, but the spatial coordination of these events is poorly understood. The Ccr4-Not complex initiates mRNA ...Control of messenger RNA (mRNA) decay rate is intimately connected to translation elongation, but the spatial coordination of these events is poorly understood. The Ccr4-Not complex initiates mRNA decay through deadenylation and activation of decapping. We used a combination of cryo-electron microscopy, ribosome profiling, and mRNA stability assays to examine the recruitment of Ccr4-Not to the ribosome via specific interaction of the Not5 subunit with the ribosomal E-site in This interaction occurred when the ribosome lacked accommodated A-site transfer RNA, indicative of low codon optimality. Loss of the interaction resulted in the inability of the mRNA degradation machinery to sense codon optimality. Our findings elucidate a physical link between the Ccr4-Not complex and the ribosome and provide mechanistic insight into the coupling of decoding efficiency with mRNA stability. |
 External links External links |  Science / Science /  PubMed:32299921 / PubMed:32299921 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.8 - 3.1 Å |
| Structure data | EMDB-10431, PDB-6tb3: EMDB-10537, PDB-6tnu: |
| Chemicals |  ChemComp-MG:  ChemComp-ZN:  ChemComp-SPD: 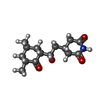 ChemComp-3HE: |
| Source |
|
 Keywords Keywords | TRANSLATION / Not5 / CCR4-NOT complex / translation control / eIF5A |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers








