+Search query
-Structure paper
| Title | Nucleotide-dependent conformational changes in the N-Ethylmaleimide Sensitive Factor (NSF) and their potential role in SNARE complex disassembly. |
|---|---|
| Journal, issue, pages | J Struct Biol, Vol. 177, Issue 2, Page 335-343, Year 2012 |
| Publish date | Jan 5, 2012 |
 Authors Authors | Arne Moeller / Chunxia Zhao / Michael G Fried / Elizabeth M Wilson-Kubalek / Bridget Carragher / Sidney W Whiteheart /  |
| PubMed Abstract | Homohexameric, N-Ethylmaleimide Sensitive Factor (NSF) disassembles Soluble NSF Attachment Protein Receptor (SNARE) complexes after membrane fusion, an essential step in vesicular trafficking. NSF ...Homohexameric, N-Ethylmaleimide Sensitive Factor (NSF) disassembles Soluble NSF Attachment Protein Receptor (SNARE) complexes after membrane fusion, an essential step in vesicular trafficking. NSF contains three domains (NSF-N, NSF-D1, and NSF-D2), each contributing to activity. We combined electron microscopic (EM) analysis, analytical ultracentrifugation (AU) and functional mutagenesis to visualize NSF's ATPase cycle. 3D density maps show that NSF-D2 remains stable, whereas NSF-N undergoes large conformational changes. NSF-Ns splay out perpendicular to the ADP-bound hexamer and twist upwards upon ATP binding, producing a more compact structure. These conformations were confirmed by hydrodynamic, AU measurements: NSF-ATP sediments faster with a lower frictional ratio (f/f(0)). Hydrodynamic analyses of NSF mutants, with specific functional defects, define the structures underlying these conformational changes. Mapping mutations onto our 3D models allows interpretation of the domain movement and suggests a mechanism for NSF binding to and disassembly of SNARE complexes. |
 External links External links |  J Struct Biol / J Struct Biol /  PubMed:22245547 / PubMed:22245547 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 16.0 - 20.0 Å |
| Structure data |  EMDB-2039: 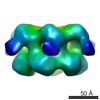 EMDB-2040:  EMDB-2041: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




