+Search query
-Structure paper
| Title | A General Small-Angle X-ray Scattering-Based Screening Protocol Validated for Protein-RNA Interactions. |
|---|---|
| Journal, issue, pages | ACS Comb Sci, Vol. 20, Issue 4, Page 197-202, Year 2018 |
| Publish date | Apr 9, 2018 |
 Authors Authors | Po-Chia Chen / Pawel Masiewicz / Vladimir Rybin / Dmitri Svergun / Janosch Hennig /  |
| PubMed Abstract | We present a screening protocol utilizing small-angle X-ray scattering (SAXS) to obtain structural information on biomolecular interactions independent of prior knowledge, so as to complement ...We present a screening protocol utilizing small-angle X-ray scattering (SAXS) to obtain structural information on biomolecular interactions independent of prior knowledge, so as to complement affinity-based screening and provide leads for further exploration. This protocol categorizes ligand titrations by computing pairwise agreement between curves, and separately estimates affinities by quantifying complex formation as a departure from the linear sum properties of solution SAXS. The protocol is validated by sparse sequence search around the native poly uridine RNA motifs of the two-RRM domain Sex-lethal protein (Sxl). The screening of 35 RNA motifs between 4 to 10 nucleotides reveals a strong variation of resulting complexes, revealed to be preference-switching between 1:1 and 2:2 binding stoichiometries upon addition of structural modeling. Validation of select sequences in isothermal calorimetry and NMR titration retrieves domain-specific roles and function of a guanine anchor. These findings reinforce the suitability of SAXS as a complement in lead identification. |
 External links External links |  ACS Comb Sci / ACS Comb Sci /  PubMed:29553252 PubMed:29553252 |
| Methods | SAS (X-ray synchrotron) |
| Structure data |  SASDD25: 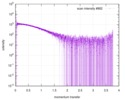 SASDDX4: Wild-type Protein sex-lethal (Protein sex-lethal, Sxl)  SASDDY4:  SASDDZ4: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




