+Search query
-Structure paper
| Title | Cryo-EM of native membranes reveals an intimate connection between the Krebs cycle and aerobic respiration in mycobacteria. |
|---|---|
| Journal, issue, pages | Proc Natl Acad Sci U S A, Vol. 122, Issue 8, Page e2423761122, Year 2025 |
| Publish date | Feb 25, 2025 |
 Authors Authors | Justin M Di Trani / Jiacheng Yu / Gautier M Courbon / Ana Paula Lobez Rodriguez / Chen-Yi Cheung / Yingke Liang / Claire E Coupland / Stephanie A Bueler / Gregory M Cook / Peter Brzezinski / John L Rubinstein /     |
| PubMed Abstract | To investigate the structure of the mycobacterial oxidative phosphorylation machinery, we prepared inverted membrane vesicles from , enriched for vesicles containing complexes of interest, and imaged ...To investigate the structure of the mycobacterial oxidative phosphorylation machinery, we prepared inverted membrane vesicles from , enriched for vesicles containing complexes of interest, and imaged the vesicles with electron cryomicroscopy. We show that this analysis allows determination of the structure of both mycobacterial ATP synthase and the supercomplex of respiratory complexes III and IV in their native membrane. The latter structure reveals that the enzyme malate:quinone oxidoreductase (Mqo) physically associates with the respiratory supercomplex, an interaction that is lost on extraction of the proteins from the lipid bilayer. Mqo catalyzes an essential reaction in the Krebs cycle, and in vivo survival of mycobacterial pathogens is compromised when its activity is absent. We show with high-speed spectroscopy that the Mqo:supercomplex interaction enables rapid electron transfer from malate to the supercomplex. Further, the respiratory supercomplex is necessary for malate-driven, but not NADH-driven, electron transport chain activity and oxygen consumption. Together, these findings indicate a connection between the Krebs cycle and aerobic respiration that directs electrons along a single branch of the mycobacterial electron transport chain. |
 External links External links |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:39969994 / PubMed:39969994 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.2 - 9.0 Å |
| Structure data | EMDB-46995, PDB-9dm1:  EMDB-47123: Cryo-EM structure of Mycobacterium smegmatis ATP synthase rotational state 1 in inverted membrane vesicles - F1 region  EMDB-47124: Cryo-EM structure of Mycobacterium smegmatis ATP synthase rotational state 1 in inverted membrane vesicles |
| Chemicals |  ChemComp-CU:  ChemComp-HEA:  ChemComp-CDL:  ChemComp-HEM:  ChemComp-MQ9: 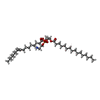 ChemComp-9Y0:  ChemComp-PLM:  ChemComp-9XX:  ChemComp-HEC:  ChemComp-9YF:  ChemComp-FES: |
| Source |
|
 Keywords Keywords | MEMBRANE PROTEIN / Electron transport chain / mycobacterial CIII2CIV2 supercomplex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers





 mycolicibacterium smegmatis mc2 155 (bacteria)
mycolicibacterium smegmatis mc2 155 (bacteria)